Project Management With RStudio
Last updated on 2025-03-19 | Edit this page
Overview
Questions
- How can I manage my projects in R?
Objectives
- Create self-contained projects in RStudio
Introduction
The scientific process is naturally incremental, and many projects start life as random notes, some code, then a manuscript, and eventually everything is a bit mixed together. Organising a project involving spatial data is no different from any other data analysis project, although you may require more disk space than usual.
Managing your projects in a reproducible fashion doesn’t just make your science reproducible, it makes your life easier.
— Vince Buffalo (@vsbuffalo) April 15, 2013
Most people tend to organize their projects like this:
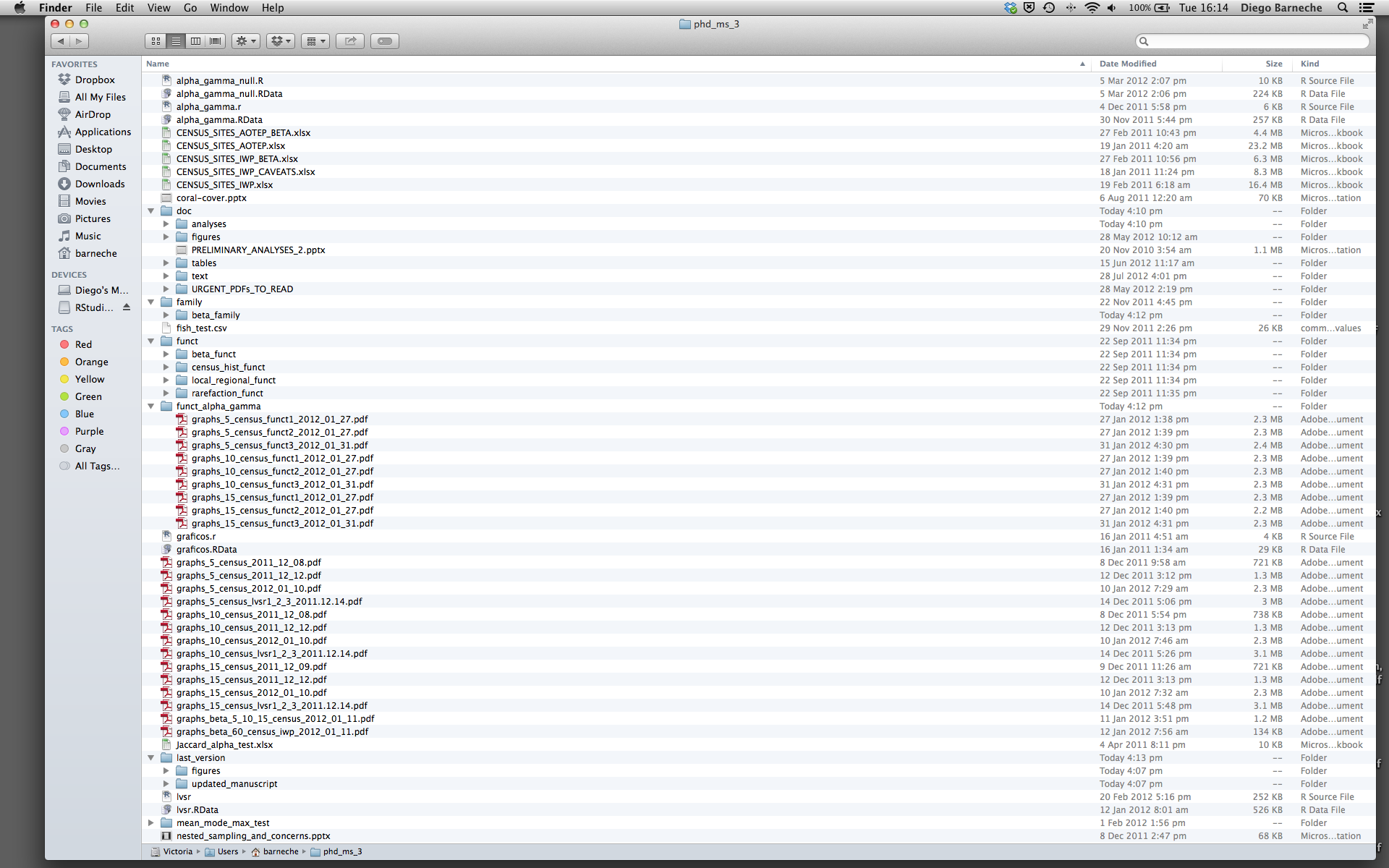
There are many reasons why we should ALWAYS avoid this:
- It is really hard to tell which version of your data is the original and which is the modified;
- It gets really messy because it mixes files with various extensions together;
- It probably takes you a lot of time to actually find things, and relate the correct figures to the exact code that has been used to generate it;
A good project layout will ultimately make your life easier:
- It will help ensure the integrity of your data;
- It makes it simpler to share your code with someone else (a lab-mate, collaborator, or supervisor);
- It allows you to easily upload your code with your manuscript submission;
- It makes it easier to pick the project back up after a break.
A possible solution
Fortunately, there are tools and packages which can help you manage your work effectively.
One of the most powerful and useful aspects of RStudio is its project management functionality. We’ll be using this today to create a self-contained, reproducible project.
Challenge: Creating a self-contained project
We’re going to create a new project in RStudio:
- Click the “File” menu button, then “New Project”.
- Click “New Directory”.
- Click “New Project”.
- Type in “r-geospatial” as the name of the directory.
- Click the “Create Project” button.
A key advantage of an RStudio Project is that whenever we open this
project in subsequent RStudio sessions our working directory will
always be set to the folder r-geospatial. Let’s
check our working directory by entering the following into the R
console:
R
getwd()R should return your/path/r-geospatial as the working
directory.
Best practices for project organization
Although there is no “best” way to lay out a project, there are some general principles to adhere to that will make project management easier:
Treat data as read only
This is probably the most important goal of setting up a project. Data is typically time consuming and/or expensive to collect. Working with them interactively (e.g., in Excel) where they can be modified means you are never sure of where the data came from, or how it has been modified since collection. It is therefore a good idea to treat your data as “read-only”.
Data Cleaning
In many cases your data will be “dirty”: it will need significant preprocessing to get into a format R (or any other programming language) will find useful. This task is sometimes called “data munging”. I find it useful to store these scripts in a separate folder, and create a second “read-only” data folder to hold the “cleaned” data sets.
Treat generated output as disposable
Anything generated by your scripts should be treated as disposable: it should all be able to be regenerated from your scripts.
There are lots of different ways to manage this output. I find it useful to have an output folder with different sub-directories for each separate analysis. This makes it easier later, as many of my analyses are exploratory and don’t end up being used in the final project, and some of the analyses get shared between projects.
Keep related data together
Some GIS file formats are really 3-6 files that need to be kept together and have the same name, e.g. shapefiles. It may be tempting to store those components separately, but your spatial data will be unusable if you do that.
Keep a consistent naming scheme
It is generally best to avoid renaming downloaded spatial data, so
that a clear connection is maintained with the point of truth. You may
otherwise find yourself wondering whether file_A really is
just a copy of Official_file_on_website or not.
For datasets you generate, it’s worth taking the time to come up with a naming convention that works for your project, and sticking to it. File names don’t have to be long, they just have to be long enough that you can tell what the file is about. Date generated, topic, and whether a product is intermediate or final are good bits of information to keep in a file name. For more tips on naming files, check out the slides from Jenny Bryan’s talk “Naming things” at the 2015 Reproducible Science Workshop.
Tip: Good Enough Practices for Scientific Computing
Good Enough Practices for Scientific Computing gives the following recommendations for project organization:
- Put each project in its own directory, which is named after the project.
- Put text documents associated with the project in the
docdirectory. - Put raw data and metadata in the
datadirectory, and files generated during cleanup and analysis in aresultsdirectory. - Put source for the project’s scripts and programs in the
srcdirectory, and programs brought in from elsewhere or compiled locally in thebindirectory. - Name all files to reflect their content or function.
Save the data in the data directory
Now we have a good directory structure we will now place/save our
data files in the data/ directory.
Challenge 1
1. Download each of the data files listed below (Ctrl+S, right mouse click -> “Save as”, or File -> “Save page as”)
2. Make sure the files have the following names:
nordic-data.csvnordic-data-2.csvgapminder_data.csv
3. Save the files in the data/ folder within your
project.
We will load and inspect these data later.
Challenge 2
We also want to move the data that we downloaded from the data
page into a subdirectory inside r-geospatial. If you
haven’t already downloaded the data, you can do so by clicking this
download link.
- Move the downloaded zip file to the
datadirectory. - Once the data have been moved, unzip all files.
Once you have completed moving the data across to the new folder, your data directory should look as follows:
data/
gapminder_data.csv
NEON-DS-Airborne-Remote-Sensing/
NEON-DS-Landsat-NDVI/
NEON-DS-Met-Time-Series/
NEON-DS-Site-Layout-Files/
NEON-DS-Airborne-Remote-Sensing.zip
NEON-DS-Landsat-NDVI.zip
NEON-DS-Met-Time-Series.zip
NEON-DS-Site-Layout-Files.zip
nordic-data.csv
nordic-data-2.csvStage your scripts
Creating separate R scripts or Rmarkdown documents for different stages of a project will maximise efficiency. For instance, separating data download commands into their own file means that you won’t re-download data unnecessarily.
Key Points
- Use RStudio to create and manage projects with consistent layout.
- Treat raw data as read-only.
- Treat generated output as disposable.
