Content from Introduction to R and RStudio
Last updated on 2024-07-23 | Edit this page
Overview
Questions
- Why should you use R and RStudio?
- How do you get started working in R and RStudio?
Objectives
- Understand the difference between R and RStudio
- Describe the purpose of the different RStudio panes
- Organize files and directories into R Projects
- Use the RStudio help interface to get help with R functions
- Be able to format questions to get help in the broader R community
What are R and RStudio?
R refers to a programming language as well as the software that runs R code.
RStudio is a software interface
that can make it easier to write R scripts and interact with the R
software. It’s a very popular platform, and RStudio also maintains the
tidyverse series
of packages we will use in this lesson.
Why learn R?
You’re working on a project when your advisor suggests that you begin working with one of their long-time collaborators. According to your advisor, this collaborator is very talented, but only speaks a language that you don’t know. Your advisor assures you that this is ok, the collaborator won’t judge you for starting to learn the language, and will happily answer your questions. However, the collaborator is also quite pedantic. While they don’t mind that you don’t speak their language fluently yet, they are always going to answer you quite literally.
You decide to reach out to the collaborator. You find that they email you back very quickly, almost immediately most of the time. Since you’re just learning their language, you often make mistakes. Sometimes, they tell you that you’ve made a grammatical error or warn you that what you asked for doesn’t make a lot of sense. Sometimes these warnings are difficult to understand, because you don’t really have a grasp of the underlying grammar. Sometimes you get an answer back, with no warnings, but you realize that it doesn’t make sense, because what you asked for isn’t quite what you wanted. Since this collaborator responds almost immediately, without tiring, you can quickly reformulate your question and send it again.
In this way, you begin to learn the language your collaborator speaks, as well as the particular way they think about your work. Eventually, the two of you develop a good working relationship, where you understand how to ask them questions effectively, and how to work through any issues in communication that might arise.
This collaborator’s name is R.
When you send commands to R, you get a response back. Sometimes, when you make mistakes, you will get back a nice, informative error message or warning. However, sometimes the warnings seem to reference a much “deeper” level of R than you’re familiar with. Or, even worse, you may get the wrong answer with no warning because the command you sent is perfectly valid, but isn’t what you actually want. While you may first have some success working with R by memorizing certain commands or reusing other scripts, this is akin to using a collection of tourist phrases or pre-written statements when having a conversation. You might make a mistake (like getting directions to the library when you need a bathroom), and you are going to be limited in your flexibility (like furiously paging through a tourist guide looking for the term for “thrift store”).
This is all to say that we are going to spend a bit of time digging
into some of the more fundamental aspects of the R language, and these
concepts may not feel as immediately useful as, say, learning to make
plots with ggplot2. However, learning these more
fundamental concepts will help you develop an understanding of how R
thinks about data and code, how to interpret error messages, and how to
flexibly expand your skills to new situations.
R does not involve lots of pointing and clicking, and that’s a good thing
Since R is a programming language, the results of your analysis do not rely on remembering a succession of pointing and clicking, but instead on a series of written commands, and that’s a good thing! So, if you want to redo your analysis because you collected more data, you don’t have to remember which button you clicked in which order to obtain your results; you just have to run your script again.
Working with scripts makes the steps you used in your analysis clear, and the code you write can be inspected by someone else who can give you feedback and spot mistakes.
Working with scripts forces you to have a deeper understanding of what you are doing, and facilitates your learning and comprehension of the methods you use.
R code is great for reproducibility
Reproducibility is when someone else (including your future self) can obtain the same results from the same dataset when using the same analysis.
R integrates with other tools to generate manuscripts from your code. If you collect more data, or fix a mistake in your dataset, the figures and the statistical tests in your manuscript are updated automatically.
An increasing number of journals and funding agencies expect analyses to be reproducible, so knowing R will give you an edge with these requirements.
R is interdisciplinary and extensible
With tens of thousands of packages that can be installed to extend its capabilities, R provides a framework that allows you to combine statistical approaches from many scientific disciplines to best suit the analytical framework you need to analyze your data. For instance, R has packages for image analysis, GIS, time series, population genetics, and a lot more.
R works on data of all shapes and sizes
The skills you learn with R scale easily with the size of your dataset. Whether your dataset has hundreds or millions of lines, it won’t make much difference to you.
R is designed for data analysis. It comes with special data structures and data types that make handling of missing data and statistical factors convenient.
R can read data from many different file types, including geospatial data, and connect to local and remote databases.
R produces high-quality graphics
R has well-developed plotting capabilities, and the
ggplot2 package is one of, if not the most powerful pieces
of plotting software available today. We will begin learning to use
ggplot2 in the next episode.
R has a large and welcoming community
Thousands of people use R daily. Many of them are willing to help you through mailing lists and websites such as Stack Overflow, or on the RStudio community.
Since R is very popular among researchers, most of the help communities and learning materials are aimed towards other researchers. Python is a similar language to R, and can accomplish many of the same tasks, but is widely used by software developers and software engineers, so Python resources and communities are not as oriented towards researchers.
Navigating RStudio
We will use the RStudio integrated development environment (IDE) to write code into scripts, run code in R, navigate files on our computer, inspect objects we create in R, and look at the plots we make. RStudio has many other features that can help with things like version control, developing R packages, and writing Shiny apps, but we won’t cover those in the workshop.
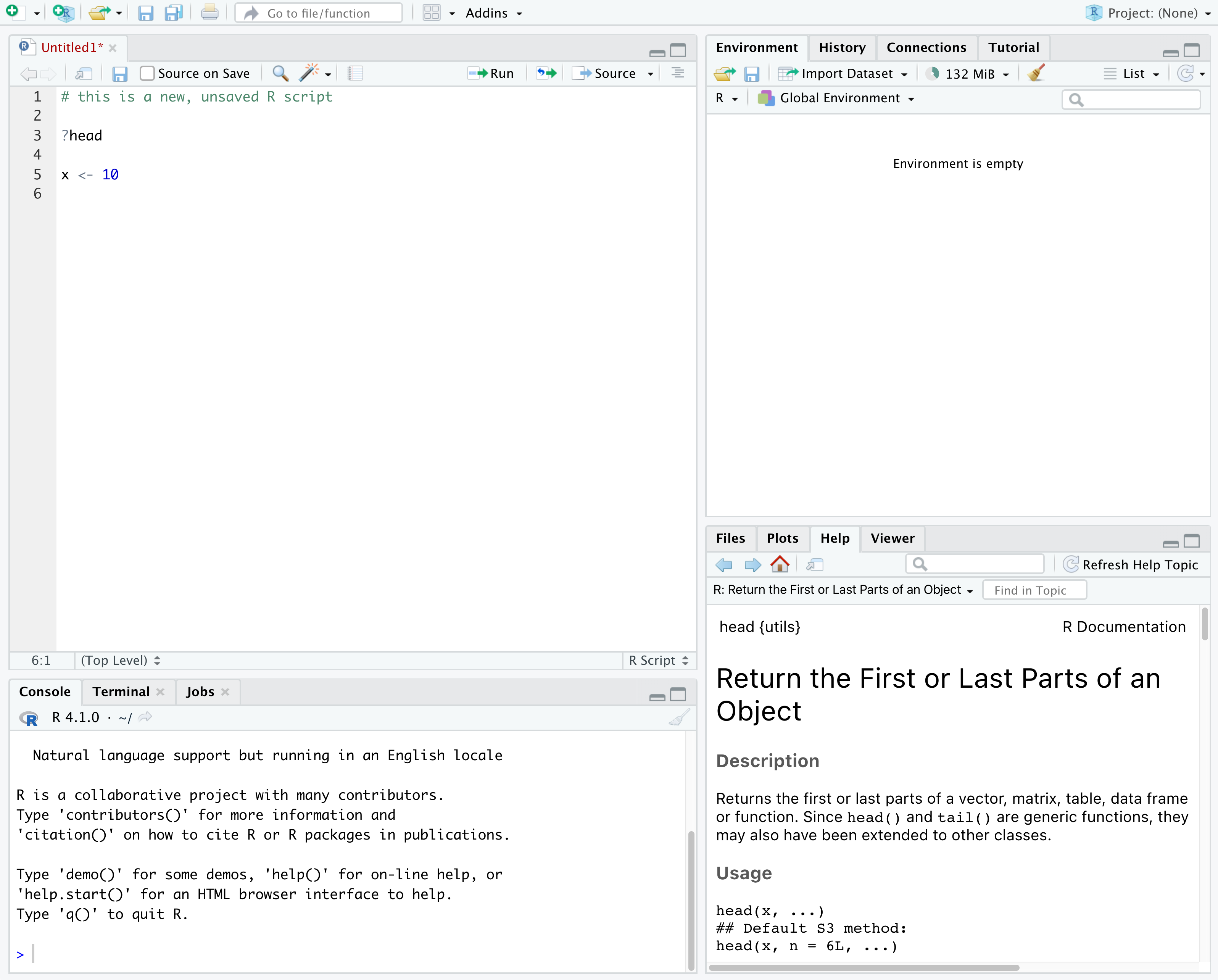
In the above screenshot, we can see 4 “panes” in the default layout:
- Top-Left: the Source pane that displays scripts and
other files.
- If you only have 3 panes, and the Console pane is in the top left, press Shift+Cmd+N (Mac) or Shift+Ctrl+N (Windows) to open a blank R script, which should make the Source pane appear.
- Top-Right: the Environment/History pane, which
shows all the objects in your current R session (Environment) and your
command history (History)
- there are some other tabs here, including Connections, Build, Tutorial, and possibly Git
- we won’t cover any of the other tabs, but RStudio has lots of other useful features
- Bottom-Left: the Console pane, where you can
interact directly with an R console, which interprets R commands and
prints the results
- There are also tabs for Terminal and Jobs
- Bottom-Right: the Files/Plots/Help/Viewer pane to navigate files or view plots and help pages
You can customize the layout of these panes, as well as many settings such as RStudio color scheme, font, and even keyboard shortcuts. You can access these settings by going to the menu bar, then clicking on Tools → Global Options.
RStudio puts most of the things you need to work in R into a single window, and also includes features like keyboard shortcuts, autocompletion of code, and syntax highlighting (different types of code are colored differently, making it easier to navigate your code).
Getting set up in RStudio
It is a good practice to organize your projects into self-contained folders right from the start, so we will start building that habit now. A well-organized project is easier to navigate, more reproducible, and easier to share with others. Your project should start with a top-level folder that contains everything necessary for the project, including data, scripts, and images, all organized into sub-folders.
RStudio provides a “Projects” feature that can make it easier to work on individual projects in R. We will create a project that we will keep everything for this workshop.
- Start RStudio (you should see a view similar to the screenshot above).
- In the top right, you will see a blue 3D cube and the words “Project: (None)”. Click on this icon.
- Click New Project from the dropdown menu.
- Click New Directory, then New Project.
- Type out a name for the project, we recommend
R-Ecology-Workshop. - Put it in a convenient location using the “Create project as a
subdirectory of:” section. We recommend your
Desktop. You can always move the project somewhere else later, because it will be self-contained. - Click Create Project and your new project will open.
Next time you open RStudio, you can click that 3D cube icon, and you will see options to open existing projects, like the one you just made.
One of the benefits to using RStudio Projects is that they
automatically set the working directory to the
top-level folder for the project. The working directory is the folder
where R is working, so it views the location of all files (including
data and scripts) as being relative to the working directory. You may
come across scripts that include something like
setwd("/Users/YourUserName/MyCoolProject"), which directly
sets a working directory. This is usually much less portable, since that
specific directory might not be found on someone else’s computer (they
probably don’t have the same username as you). Using RStudio Projects
means we don’t have to deal with manually setting the working
directory.
There are a few settings we will need to adjust to improve the reproducibility of our work. Go to your menu bar, then click Tools → Global Options to open up the Options window.
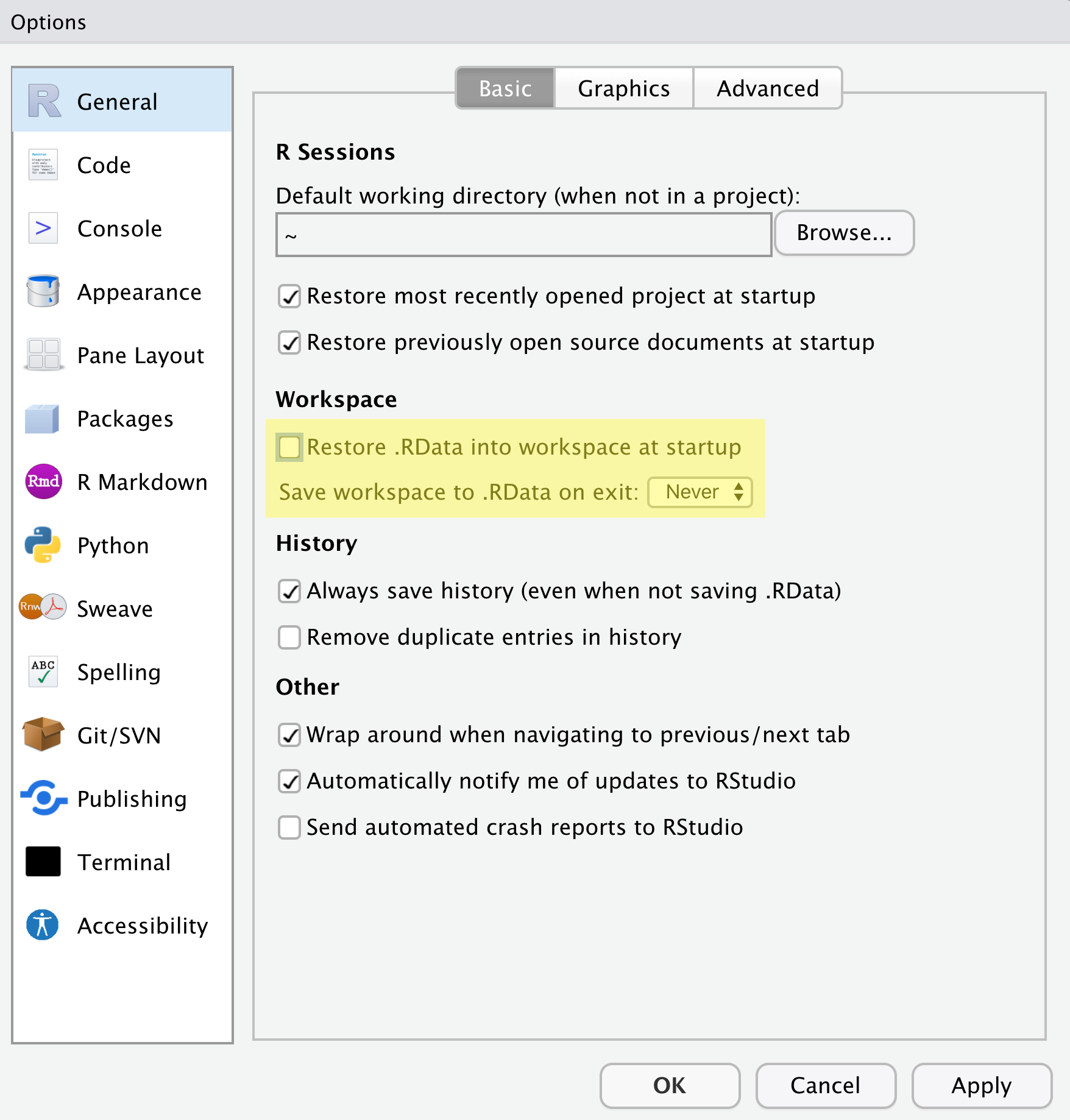
Make sure your settings match those highlighted in yellow. We don’t want RStudio to store the current status of our R session and reload it the next time we start R. This might sound convenient, but for the sake of reproducibility, we want to start with a clean, empty R session every time we work. That means that we have to record everything we do into scripts, save any data we need into files, and store outputs like images as files. We want to get used to everything we generate in a single R session being disposable. We want our scripts to be able to regenerate things we need, other than “raw materials” like data.
Organizing your project directory
Using a consistent folder structure across all your new projects will help keep a growing project organized, and make it easy to find files in the future. This is especially beneficial if you are working on multiple projects, since you will know where to look for particular kinds of files.
We will use a basic structure for this workshop, which is often a good place to start, and can be extended to meet your specific needs. Here is a diagram describing the structure:
R-Ecology-Workshop
│
└── scripts
│
└── data
│ └── cleaned
│ └── raw
│
└─── images
│
└─── documentsWithin our project folder (R-Ecology-Workshop), we first
have a scripts folder to hold any scripts we write. We also
have a data folder containing cleaned and
raw subfolders. In general, you want to keep your
raw data completely untouched, so once you put data into
that folder, you do not modify it. Instead, you read it into R, and if
you make any modifications, you write that modified file into the
cleaned folder. We also have an images folder
for plots we make, and a documents folder for any other
documents you might produce.
Let’s start making our new folders. Go to the Files
pane (bottom right), and check the current directory, highlighted in
yellow below. You should be in the directory for the project you just
made, in our case R-Ecology-Workshop. You shouldn’t see any
folders in here yet.
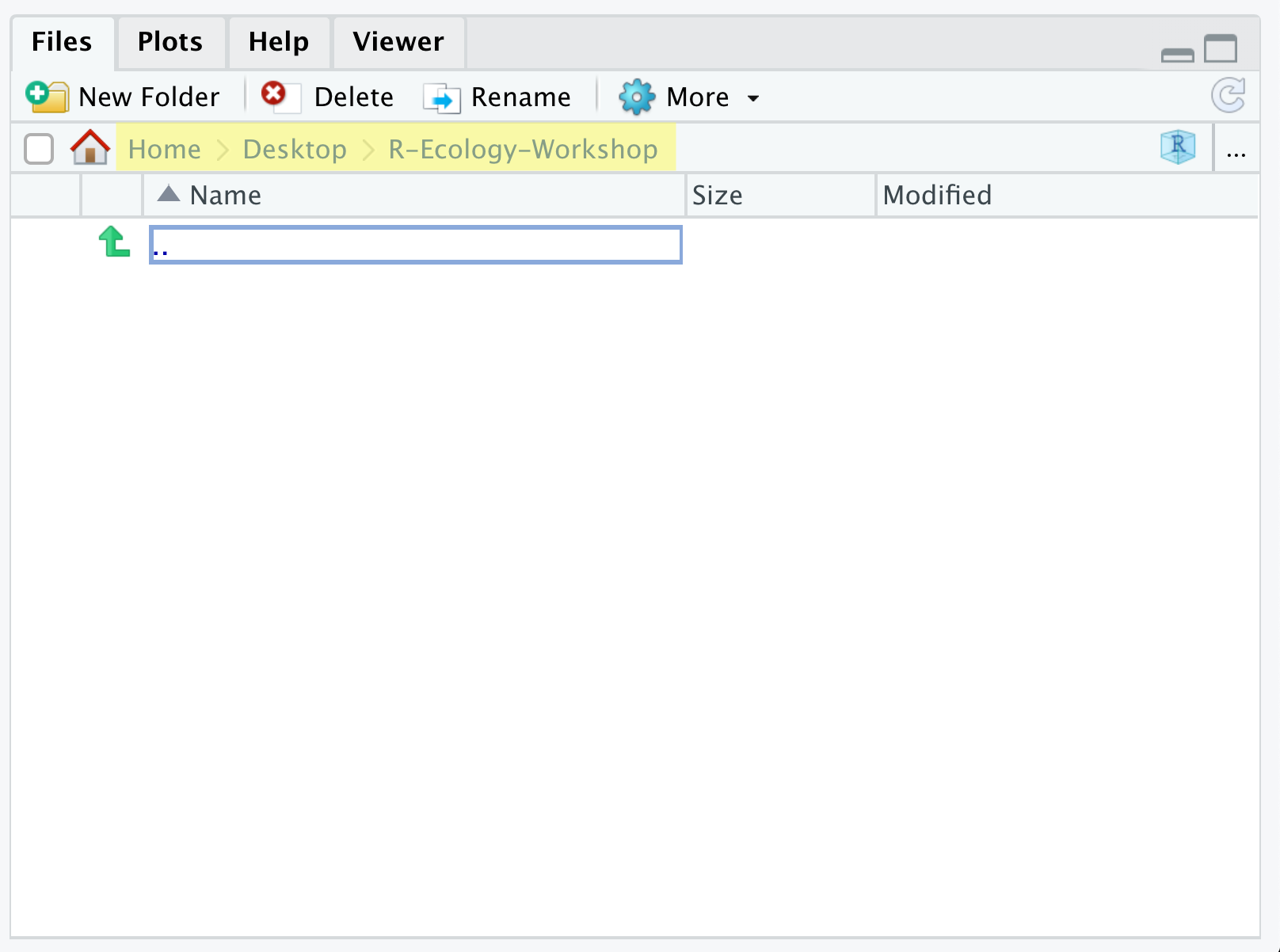
Next, click the New Folder button, and type in
scripts to generate your scripts folder. It
should appear in the Files list now. Repeat the process to make your
data, images, and documents
folders. Then, click on the data folder in the Files pane.
This will take you into the data folder, which will be
empty. Use the New Folder button to create
raw and cleaned folders. To return to the
R-Ecology-Workshop folder, click on it in the file path,
which is highlighted in yellow in the previous image. It’s worth noting
that the Files pane helps you create, find, and open
files, but moving through your files won’t change where the
working directory of your project is.
Working in R and RStudio
The basis of programming is that we write down instructions for the computer to follow, and then we tell the computer to follow those instructions. We write these instructions in the form of code, which is a common language that is understood by the computer and humans (after some practice). We call these instructions commands, and we tell the computer to follow the instructions by running (also called executing) the commands.
Console vs. script
You can run commands directly in the R console, or you can write them into an R script. It may help to think of working in the console vs. working in a script as something like cooking. The console is like making up a new recipe, but not writing anything down. You can carry out a series of steps and produce a nice, tasty dish at the end. However, because you didn’t write anything down, it’s harder to figure out exactly what you did, and in what order.
Writing a script is like taking nice notes while cooking- you can tweak and edit the recipe all you want, you can come back in 6 months and try it again, and you don’t have to try to remember what went well and what didn’t. It’s actually even easier than cooking, since you can hit one button and the computer “cooks” the whole recipe for you!
An additional benefit of scripts is that you can leave
comments for yourself or others to read. Lines that
start with # are considered comments and will not be
interpreted as R code.
Console
- The R console is where code is run/executed
- The prompt, which is the
>symbol, is where you can type commands - By pressing Enter, R will execute those commands and print the result.
- You can work here, and your history is saved in the History pane, but you can’t access it in the future
Script
- A script is a record of commands to send to R, preserved in a plain
text file with a
.Rextension - You can make a new R script by clicking
File → New File → R Script, clicking the green+button in the top left corner of RStudio, or pressing Shift+Cmd+N (Mac) or Shift+Ctrl+N (Windows). It will be unsaved, and called “Untitled1” - If you type out lines of R code in a script, you can send them to
the R console to be evaluated
- Cmd+Enter (Mac) or Ctrl+Enter (Windows) will run the line of code that your cursor is on
- If you highlight multiple lines of code, you can run all of them by pressing Cmd+Enter (Mac) or Ctrl+Enter (Windows)
- By preserving commands in a script, you can edit and rerun them quickly, save them for later, and share them with others
- You can leave comments for yourself by starting a line with a
#
Example
Let’s try running some code in the console and in a script. First,
click down in the Console pane, and type out 1+1. Hit
Enter to run the code. You should see your code echoed, and
then the value of 2 returned.
Now click into your blank script, and type out 1+1. With
your cursor on that line, hit Cmd+Enter (Mac) or
Ctrl+Enter (Windows) to run the code. You will see that your
code was sent from the script to the console, where it returned a value
of 2, just like when you ran your code directly in the
console.
Content from Data visualization with ggplot2
Last updated on 2024-07-23 | Edit this page
Overview
Questions
- How do you make plots using R?
- How do you customize and modify plots?
Objectives
- Produce scatter plots and boxplots using
ggplot2. - Represent data variables with plot components.
- Modify the scales of plot components.
- Iteratively build and modify
ggplot2plots by adding layers. - Change the appearance of existing
ggplot2plots using premade and customized themes. - Describe what faceting is and apply faceting in
ggplot2. - Save plots as image files.
Setup
We are going to be using functions from the
ggplot2 package to create visualizations
of data. Functions are predefined bits of code that automate more
complicated actions. R itself has many built-in functions, but we can
access many more by loading other packages of functions
and data into R.
If you don’t have a blank, untitled script open yet, go ahead and
open one with Shift+Cmd+N (Mac) or Shift+Ctrl+N
(Windows). Then save the file to your scripts/ folder, and
title it workshop_code.R.
Earlier, you had to install the ggplot2
package by running install.packages("ggplot2"). That
installed the package onto your computer so that R can access it. In
order to use it in our current session, we have to load
the package using the library() function.
Callout
If you do not have ggplot2 installed, you can run
install.packages("ggplot2") in the
console.
It is a good practice not to put install.packages() into
a script. This is because every time you run that whole script, the
package will be reinstalled, which is typically unnecessary. You want to
install the package to your computer once, and then load it with
library() in each script where you need to use it.
R
library(ggplot2)
Later we will learn how to read data from external files into R, but
for now we are going to use a clean and ready-to-use dataset that is
provided by the ratdat data package. To
make our dataset available, we need to load this package too.
R
library(ratdat)
The ratdat package contains data from the Portal Project, which
is a long-term dataset from Portal, Arizona, in the Chihuahuan
desert.
We will be using a dataset called complete_old, which
contains older years of survey data. Let’s try to learn a little bit
about the data. We can use a ? in front of the name of the
dataset, which will bring up the help page for the data.
R
?complete_old
Here we can read descriptions of each variable in our data.
To actually take a look at the data, we can use the
View() function to open an interactive viewer, which
behaves like a simplified version of a spreadsheet program. It’s a handy
function, but somewhat limited when trying to view large datasets.
R
View(complete_old)
If you hover over the tab for the interactive View(),
you can click the “x” that appears, which will close the tab.
We can find out more about the dataset by using the
str() function to examine the structure of
the data.
R
str(complete_old)
OUTPUT
tibble [16,878 × 13] (S3: tbl_df/tbl/data.frame)
$ record_id : int [1:16878] 1 2 3 4 5 6 7 8 9 10 ...
$ month : int [1:16878] 7 7 7 7 7 7 7 7 7 7 ...
$ day : int [1:16878] 16 16 16 16 16 16 16 16 16 16 ...
$ year : int [1:16878] 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977 ...
$ plot_id : int [1:16878] 2 3 2 7 3 1 2 1 1 6 ...
$ species_id : chr [1:16878] "NL" "NL" "DM" "DM" ...
$ sex : chr [1:16878] "M" "M" "F" "M" ...
$ hindfoot_length: int [1:16878] 32 33 37 36 35 14 NA 37 34 20 ...
$ weight : int [1:16878] NA NA NA NA NA NA NA NA NA NA ...
$ genus : chr [1:16878] "Neotoma" "Neotoma" "Dipodomys" "Dipodomys" ...
$ species : chr [1:16878] "albigula" "albigula" "merriami" "merriami" ...
$ taxa : chr [1:16878] "Rodent" "Rodent" "Rodent" "Rodent" ...
$ plot_type : chr [1:16878] "Control" "Long-term Krat Exclosure" "Control" "Rodent Exclosure" ...str() will tell us how many observations/rows (obs) and
variables/columns we have, as well as some information about each of the
variables. We see the name of a variable (such as year),
followed by the kind of variable (int for integer,
chr for character), and the first 10 entries in that
variable. We will talk more about different data types and structures
later on.
Plotting with ggplot2
ggplot2 is a powerful package that
allows you to create complex plots from tabular data (data in a table
format with rows and columns). The gg in
ggplot2 stands for “grammar of graphics”,
and the package uses consistent vocabulary to create plots of widely
varying types. Therefore, we only need small changes to our code if the
underlying data changes or we decide to make a box plot instead of a
scatter plot. This approach helps you create publication-quality plots
with minimal adjusting and tweaking.
ggplot2 is part of the
tidyverse series of packages, which tend
to like data in the “long” or “tidy” format, which means each column
represents a single variable, and each row represents a single
observation. Well-structured data will save you lots of time making
figures with ggplot2. For now, we will use
data that are already in this format. We start learning R by using
ggplot2 because it relies on concepts that
we will need when we talk about data transformation in the next
lessons.
ggplot plots are built step by step by
adding new layers, which allows for extensive flexibility and
customization of plots.
To build a plot, we will use a basic template that can be used for different types of plots:
We use the ggplot() function to create a plot. In order
to tell it what data to use, we need to specify the data
argument. An argument is an input that a function
takes, and you set arguments using the = sign.
R
ggplot(data = complete_old)

We get a blank plot because we haven’t told ggplot()
which variables we want to correspond to parts of the plot. We can
specify the “mapping” of variables to plot elements, such as x/y
coordinates, size, or shape, by using the aes() function.
We’ll also add a comment, which is any line starting with a
#. It’s a good idea to use comments to organize your code
or clarify what you are doing.
R
# adding a mapping to x and y axes
ggplot(data = complete_old, mapping = aes(x = weight, y = hindfoot_length))
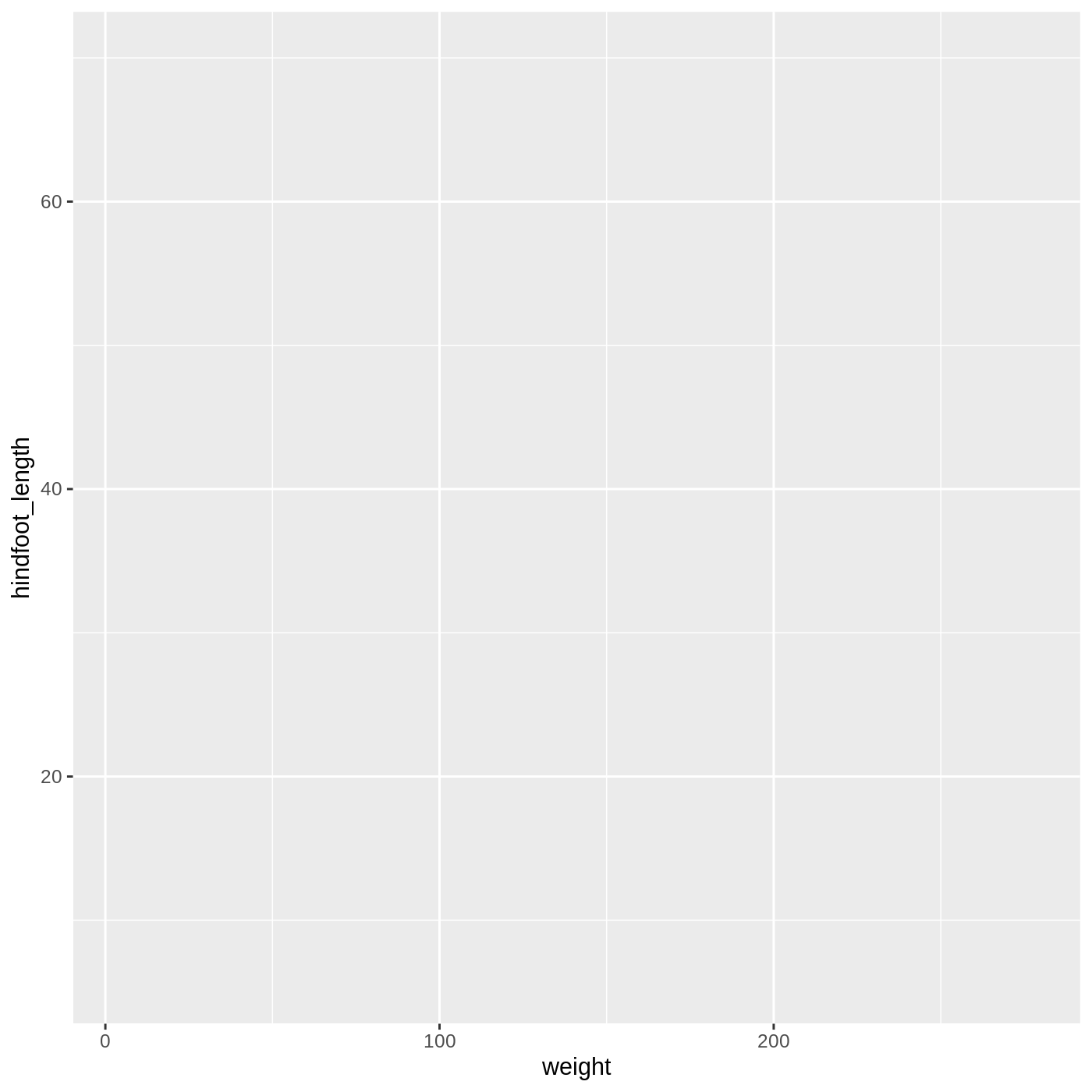
Now we’ve got a plot with x and y axes corresponding to variables
from complete_old. However, we haven’t specified how we
want the data to be displayed. We do this using geom_
functions, which specify the type of geometry we want, such
as points, lines, or bars. We can add a geom_point() layer
to our plot by using the + sign. We indent onto a new line
to make it easier to read, and we have to end the first
line with the + sign.
R
ggplot(data = complete_old, mapping = aes(x = weight, y = hindfoot_length)) +
geom_point()
WARNING
Warning: Removed 3081 rows containing missing values or values outside the scale range
(`geom_point()`).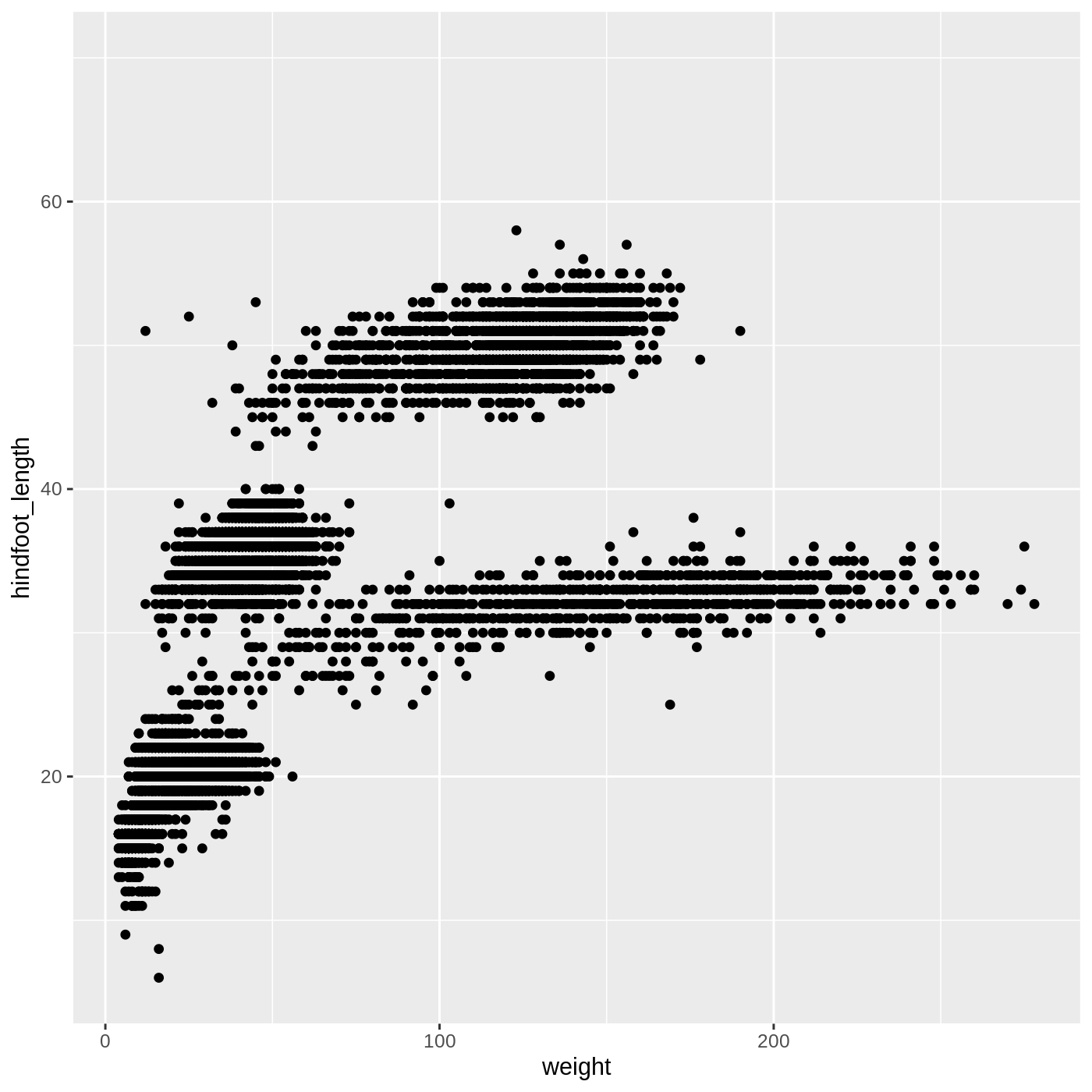
You may notice a warning that missing values were removed. If a
variable necessary to make the plot is missing from a given row of data
(in this case, hindfoot_length or weight), it
can’t be plotted. ggplot2 just uses a warning message to
let us know that some rows couldn’t be plotted.
Callout
Warning messages are one of a few ways R will communicate with you. Warnings can be thought of as a “heads up”. Nothing necessarily went wrong, but the author of that function wanted to draw your attention to something. In the above case, it’s worth knowing that some of the rows of your data were not plotted because they had missing data.
A more serious type of message is an error. Here’s an example:
R
ggplot(data = complete_old, mapping = aes(x = weight, y = hindfoot_length)) +
geom_poit()
ERROR
Error in geom_poit(): could not find function "geom_poit"As you can see, we only get the error message, with no plot, because
something has actually gone wrong. This particular error message is
fairly common, and it happened because we misspelled point
as poit. Because there is no function named
geom_poit(), R tells us it can’t find a function with that
name.
Changing aesthetics
Building ggplot plots is often an
iterative process, so we’ll continue developing the scatter plot we just
made. You may have noticed that parts of our scatter plot have many
overlapping points, making it difficult to see all the data. We can
adjust the transparency of the points using the alpha
argument, which takes a value between 0 and 1:
R
ggplot(data = complete_old, mapping = aes(x = weight, y = hindfoot_length)) +
geom_point(alpha = 0.2)
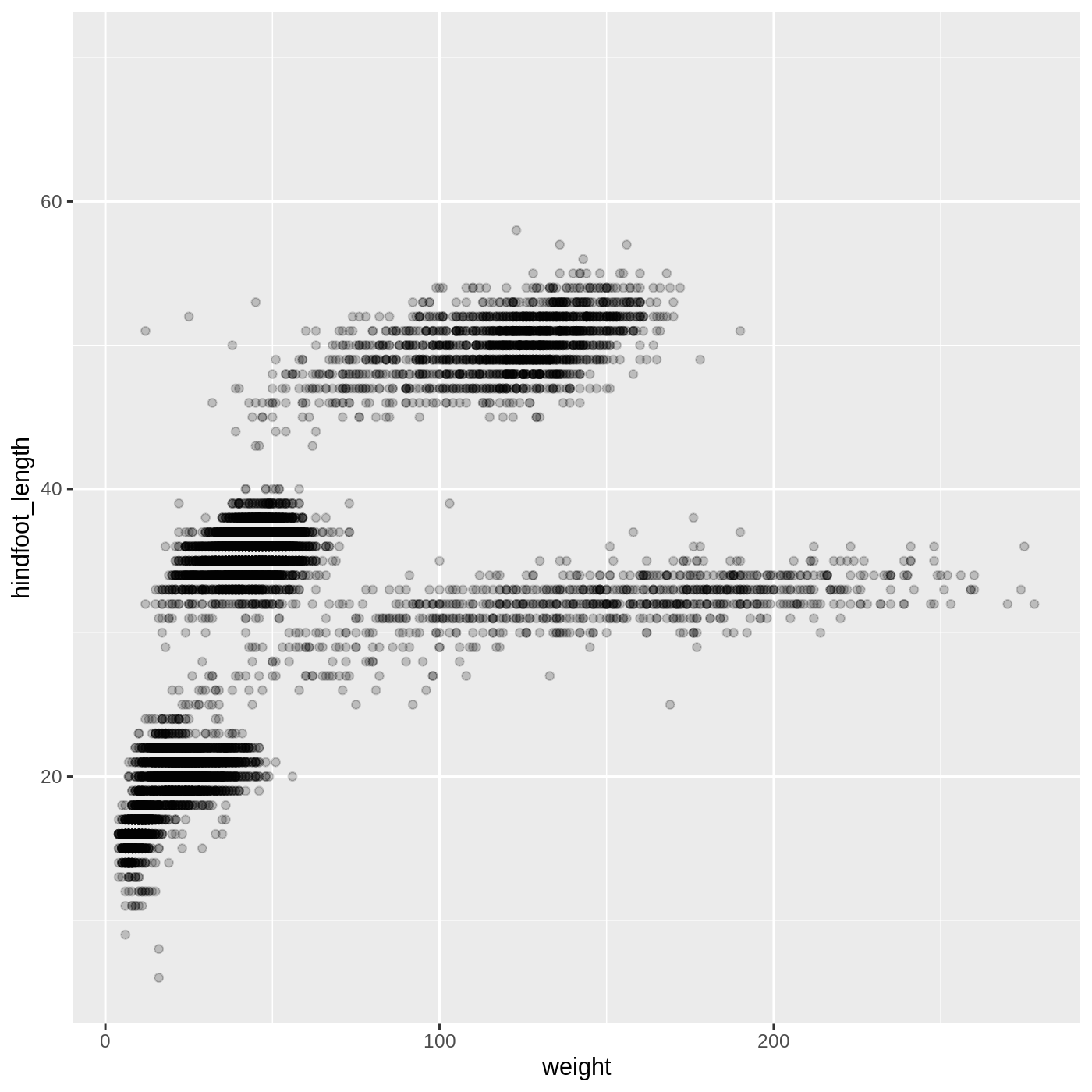
We can also change the color of the points:
R
ggplot(data = complete_old, mapping = aes(x = weight, y = hindfoot_length)) +
geom_point(alpha = 0.2, color = "blue")
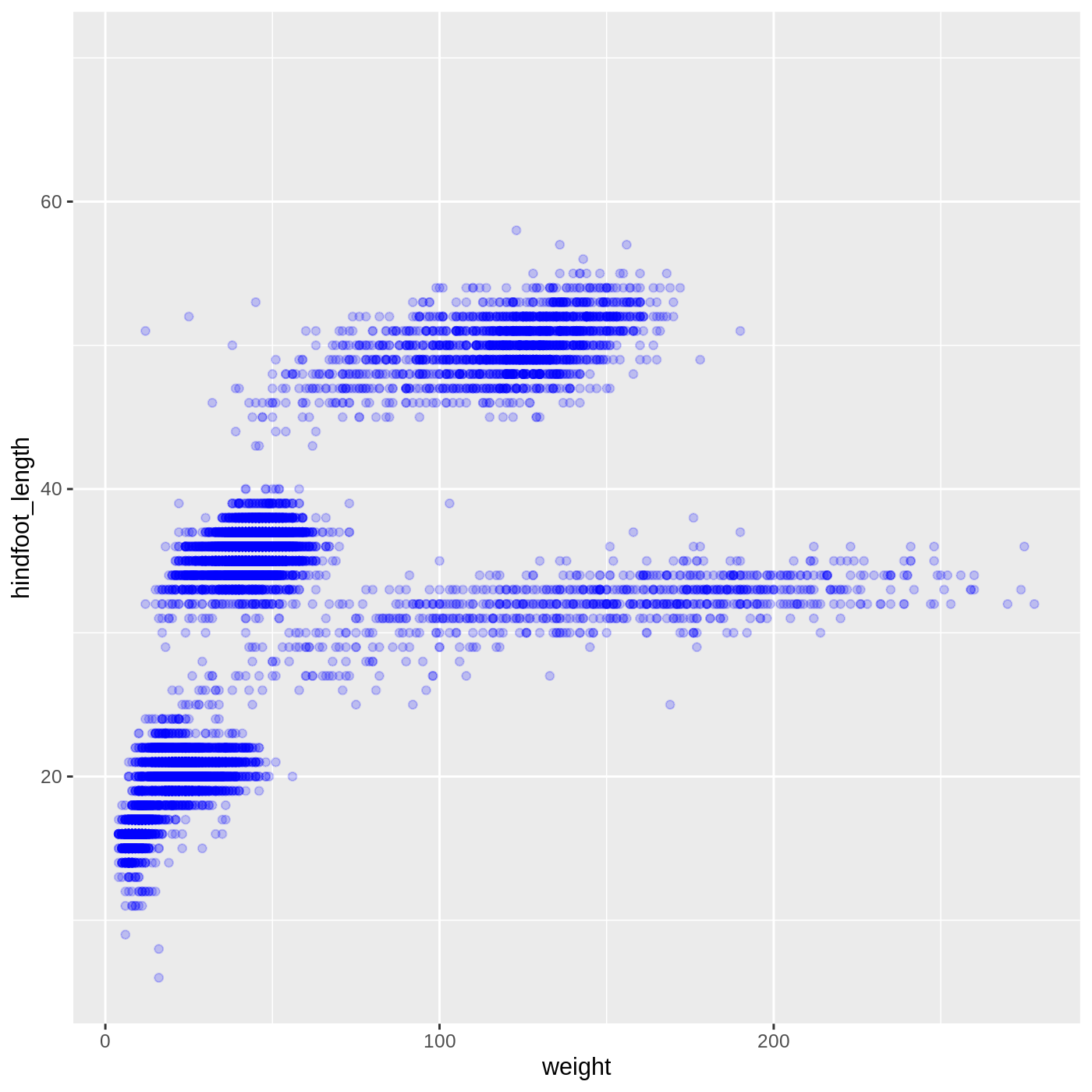
Callout
Two common issues you might run into when working in R are forgetting a closing bracket or a closing quote. Let’s take a look at what each one does.
Try running the following code:
R
ggplot(data = complete_old, mapping = aes(x = weight, y = hindfoot_length)) +
geom_point(color = "blue", alpha = 0.2You will see a + appear in your console. This is R
telling you that it expects more input in order to finish running the
code. It is missing a closing bracket to end the geom_point
function call. You can hit Esc in the console to reset
it.
Something similar will happen if you run the following code:
R
ggplot(data = complete_old, mapping = aes(x = weight, y = hindfoot_length)) +
geom_point(color = "blue, alpha = 0.2)A missing quote at the end of blue means that the rest
of the code is treated as part of the quote, which is a bit easier to
see since RStudio displays character strings in a different color.
You will get a different error message if you run the following code:
R
ggplot(data = complete_old, mapping = aes(x = weight, y = hindfoot_length)) +
geom_point(color = "blue", alpha = 0.2))This time we have an extra closing ), which R doesn’t
know what to do with. It tells you there is an unexpected
), but it doesn’t pinpoint exactly where. With enough time
working in R, you will get better at spotting mismatched brackets.
Adding another variable
Let’s try coloring our points according to the sampling plot type
(plot here refers to the physical area where rodents were sampled and
has nothing to do with making graphs). Since we’re now mapping a
variable (plot_type) to a component of the ggplot2 plot
(color), we need to put the argument inside
aes():
R
ggplot(data = complete_old, mapping = aes(x = weight, y = hindfoot_length, color = plot_type)) +
geom_point(alpha = 0.2)
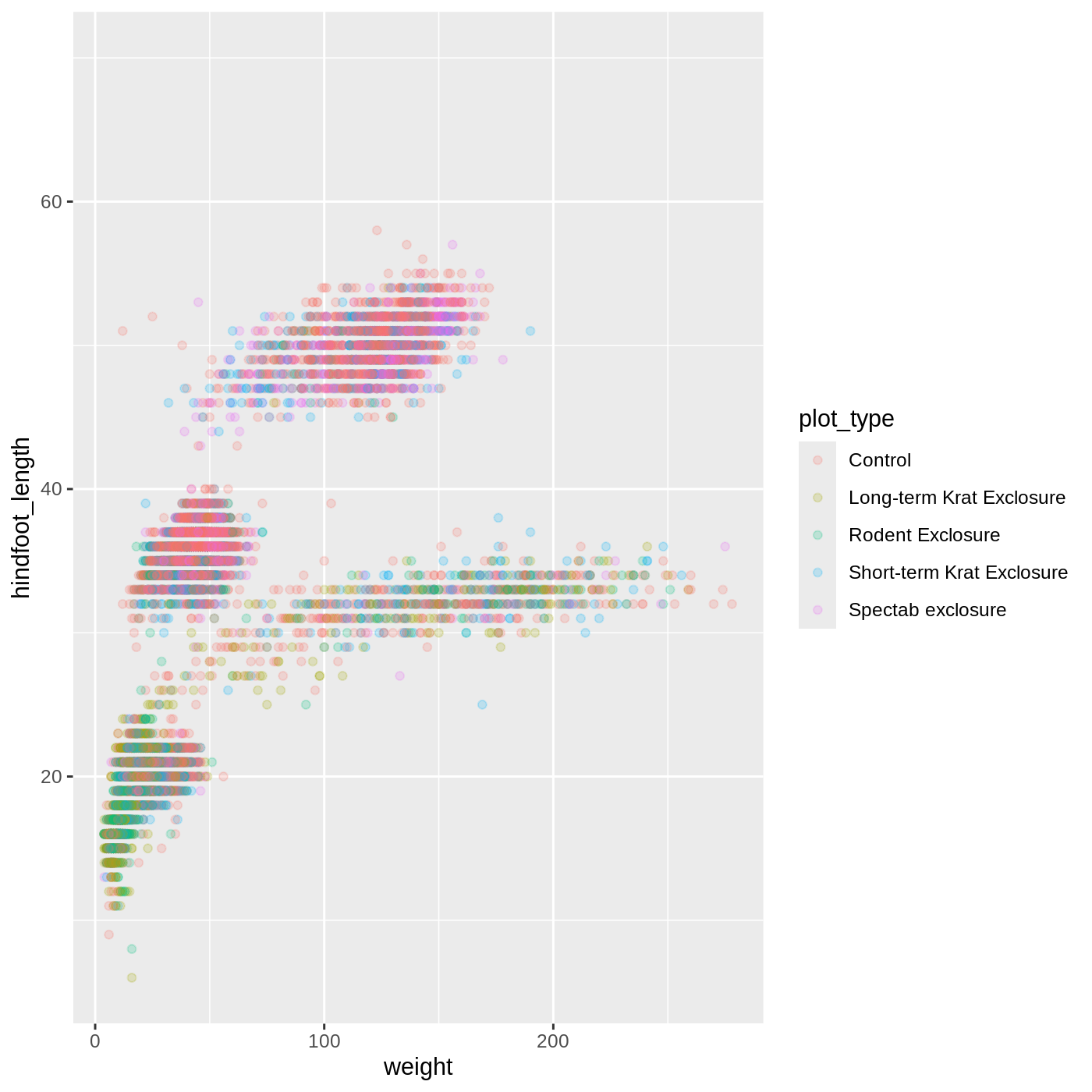
R
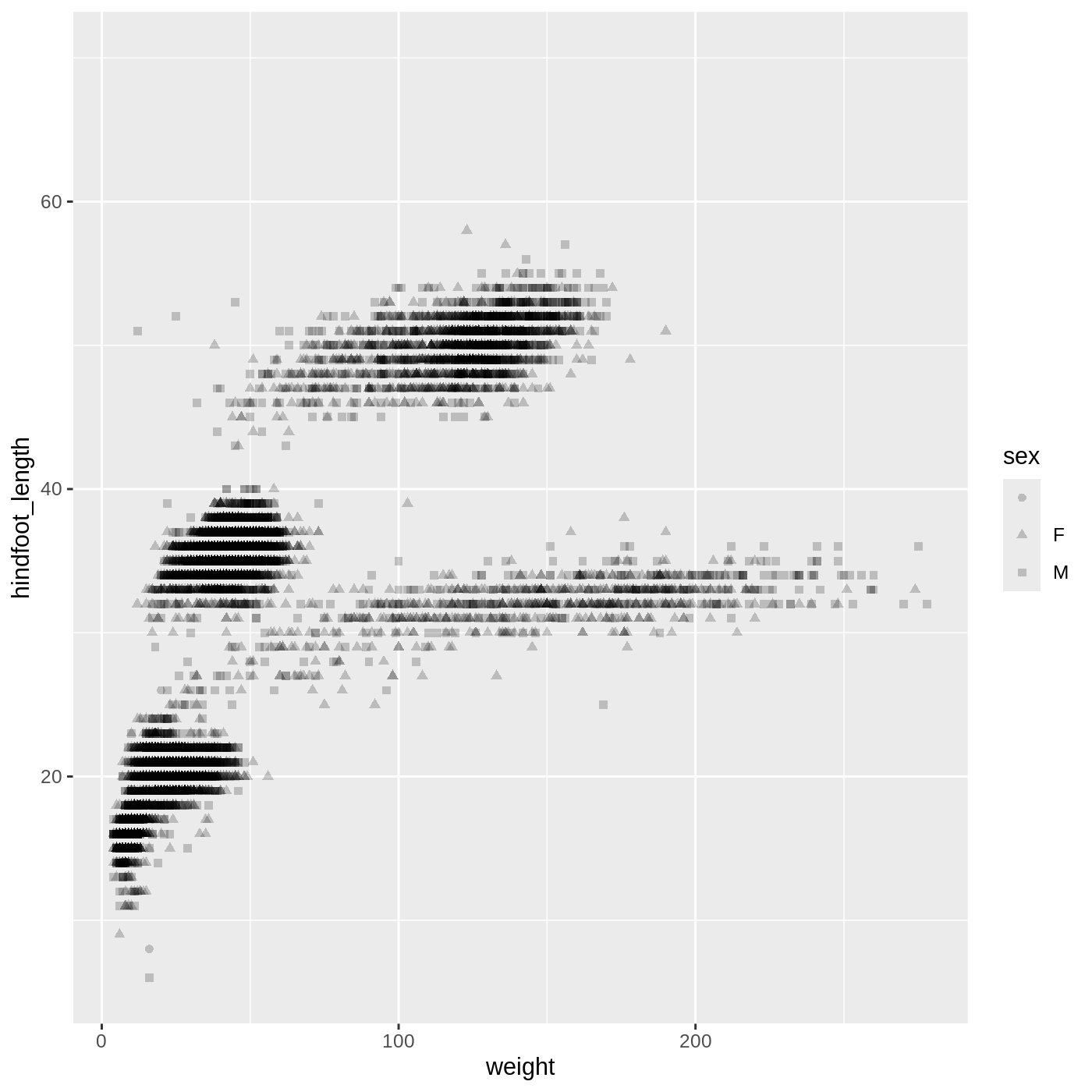
ggplot(data = complete_old,
mapping = aes(x = weight, y = hindfoot_length, shape = sex)) +
geom_point(alpha = 0.2)
R
ggplot(data = complete_old,
mapping = aes(x = weight, y = hindfoot_length, color = year)) +
geom_point(alpha = 0.2)
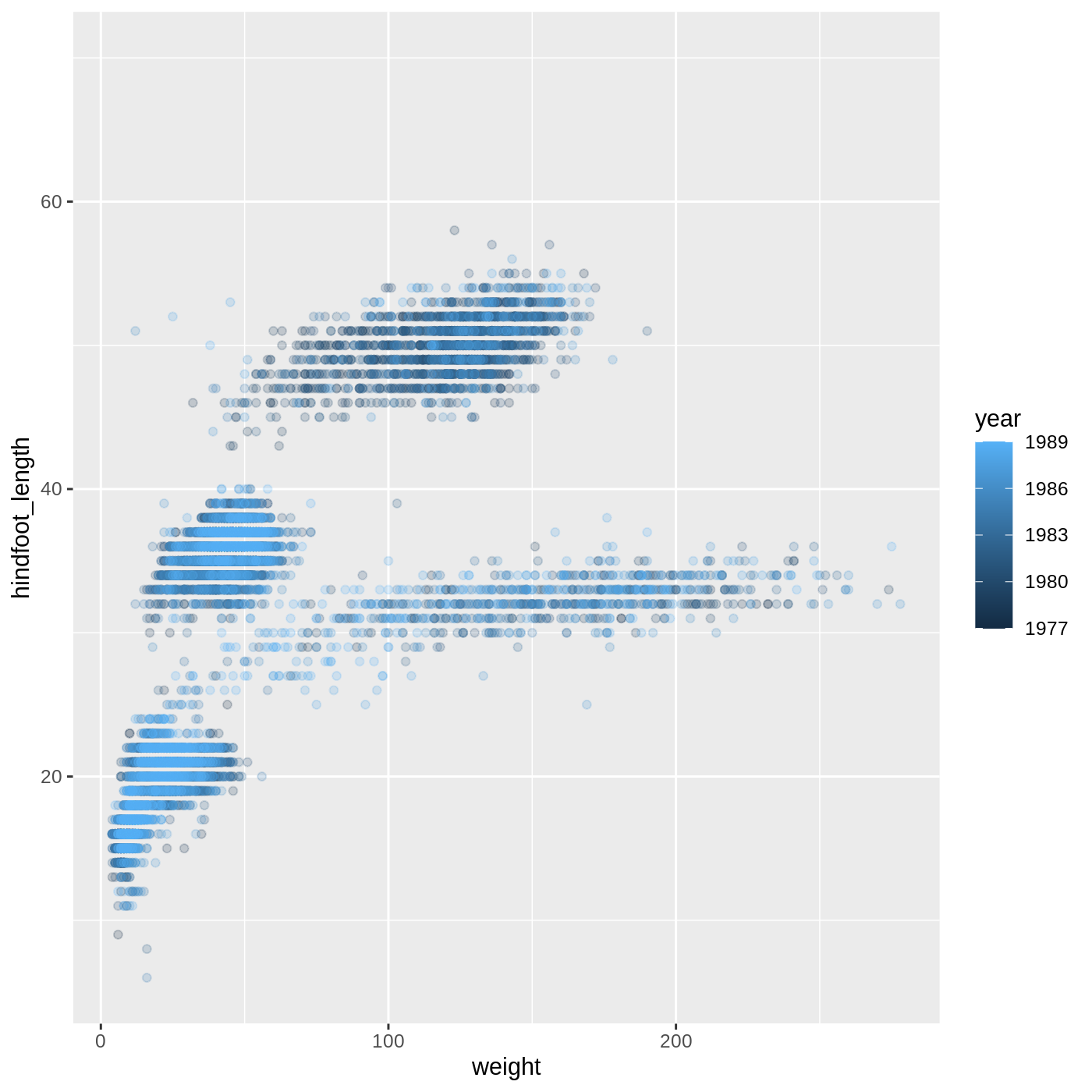
- For Part 2, the color scale is different compared to using
color = plot_typebecauseplot_typeandyearare different variable types.plot_typeis a categorical variable, soggplot2defaults to use a discrete color scale, whereasyearis a numeric variable, soggplot2uses a continuous color scale.
Changing scales
The default discrete color scale isn’t always ideal: it isn’t
friendly to viewers with colorblindness and it doesn’t translate well to
grayscale. However, ggplot2 comes with
quite a few other color scales, including the fantastic
viridis scales, which are designed to be colorblind and
grayscale friendly. We can change scales by adding scale_
functions to our plots:
R
ggplot(data = complete_old, mapping = aes(x = weight, y = hindfoot_length, color = plot_type)) +
geom_point(alpha = 0.2) +
scale_color_viridis_d()
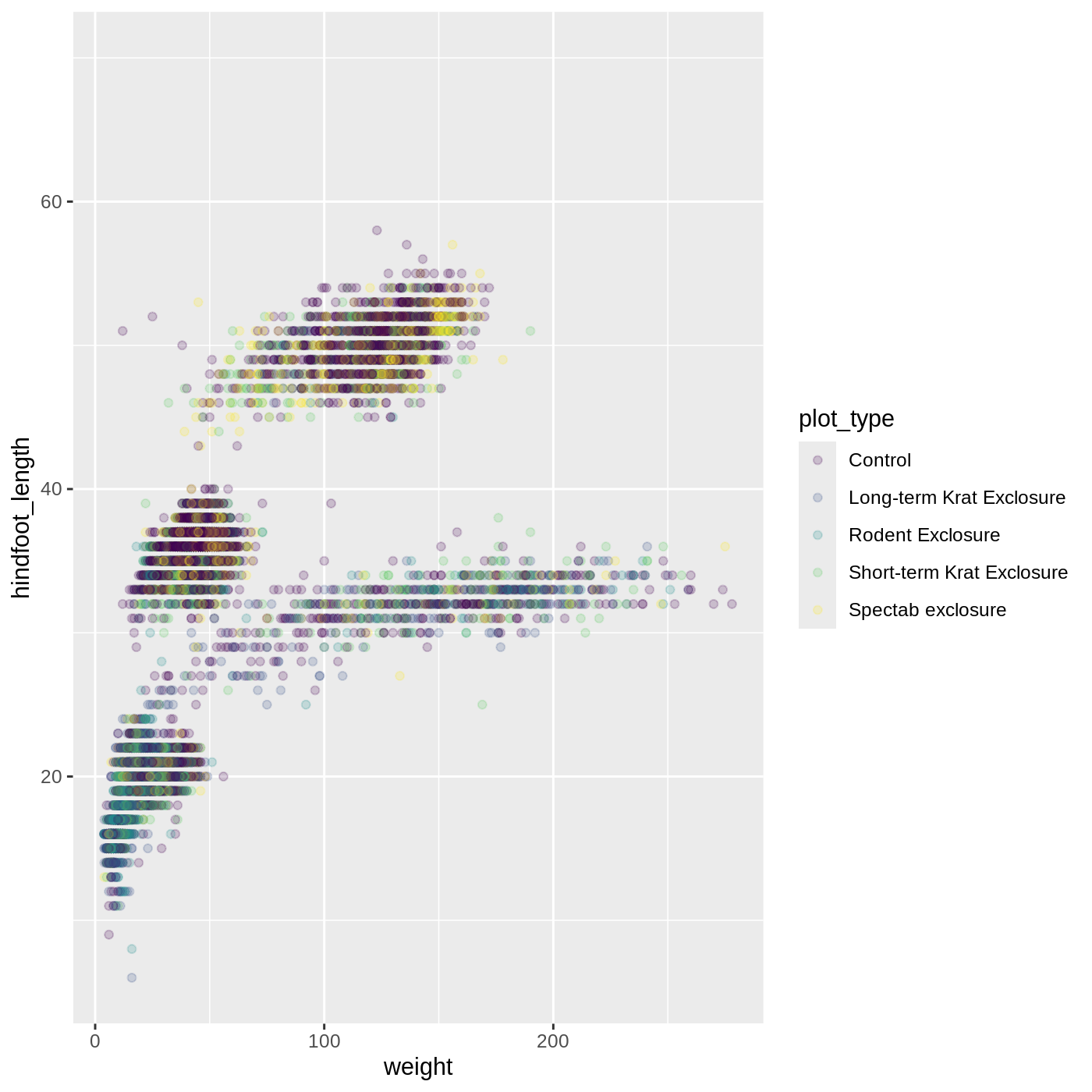
Scales don’t just apply to colors- any plot component that you put
inside aes() can be modified with scale_
functions. Just as we modified the scale used to map
plot_type to color, we can modify the way that
weight is mapped to the x axis by using the
scale_x_log10() function:
R
ggplot(data = complete_old, mapping = aes(x = weight, y = hindfoot_length, color = plot_type)) +
geom_point(alpha = 0.2) +
scale_x_log10()
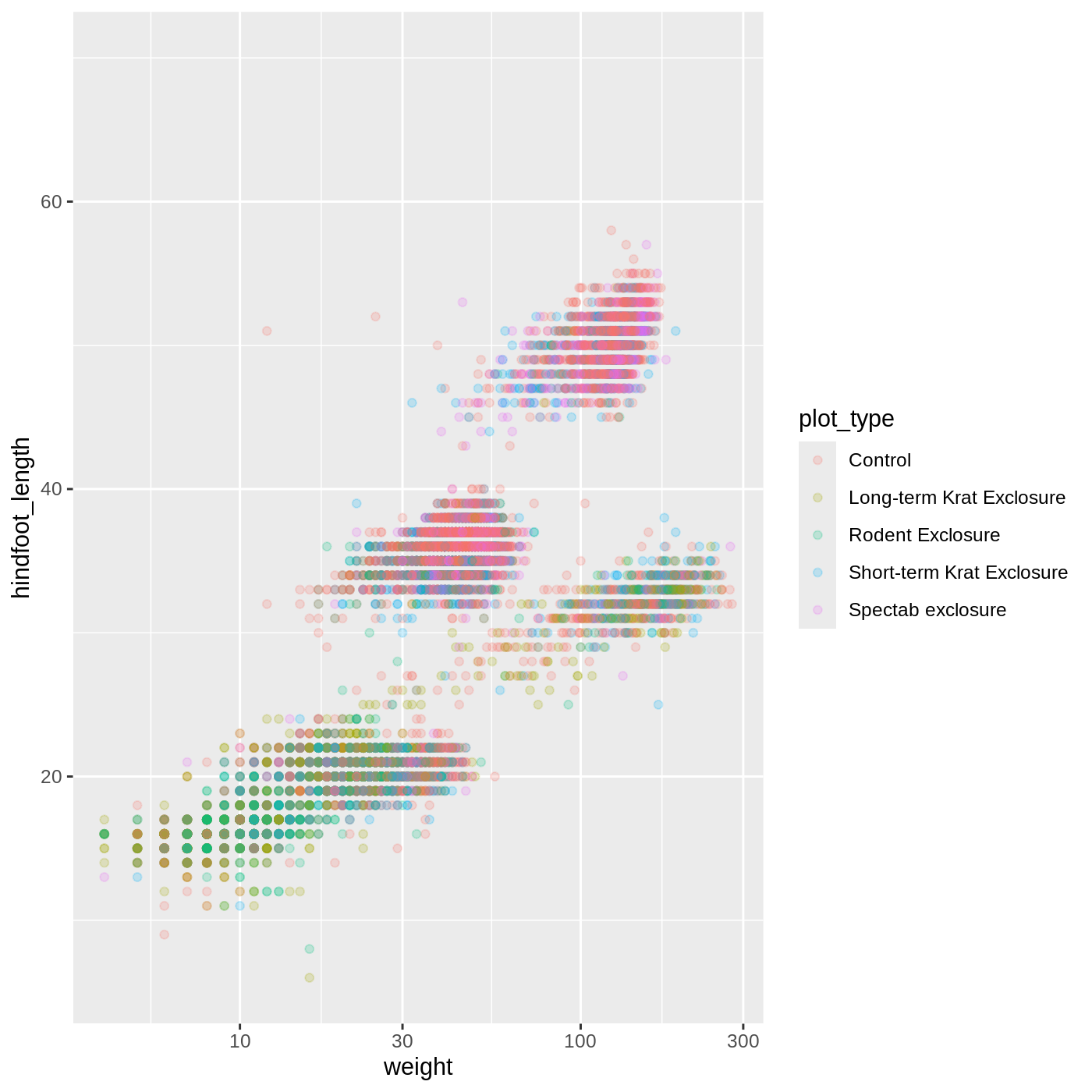
One nice thing about ggplot and the
tidyverse in general is that groups of functions that do
similar things are given similar names. Any function that modifies a
ggplot scale starts with scale_, making it
easier to search for the right function.
Boxplot
Let’s try making a different type of plot altogether. We’ll start off
with our same basic building blocks using ggplot() and
aes().
R
ggplot(data = complete_old, mapping = aes(x = plot_type, y = hindfoot_length))
This time, let’s try making a boxplot, which will have
plot_type on the x axis and hindfoot_length on
the y axis. We can do this by adding geom_boxplot() to our
ggplot():
R
ggplot(data = complete_old, mapping = aes(x = plot_type, y = hindfoot_length)) +
geom_boxplot()
WARNING
Warning: Removed 2733 rows containing non-finite outside the scale range
(`stat_boxplot()`).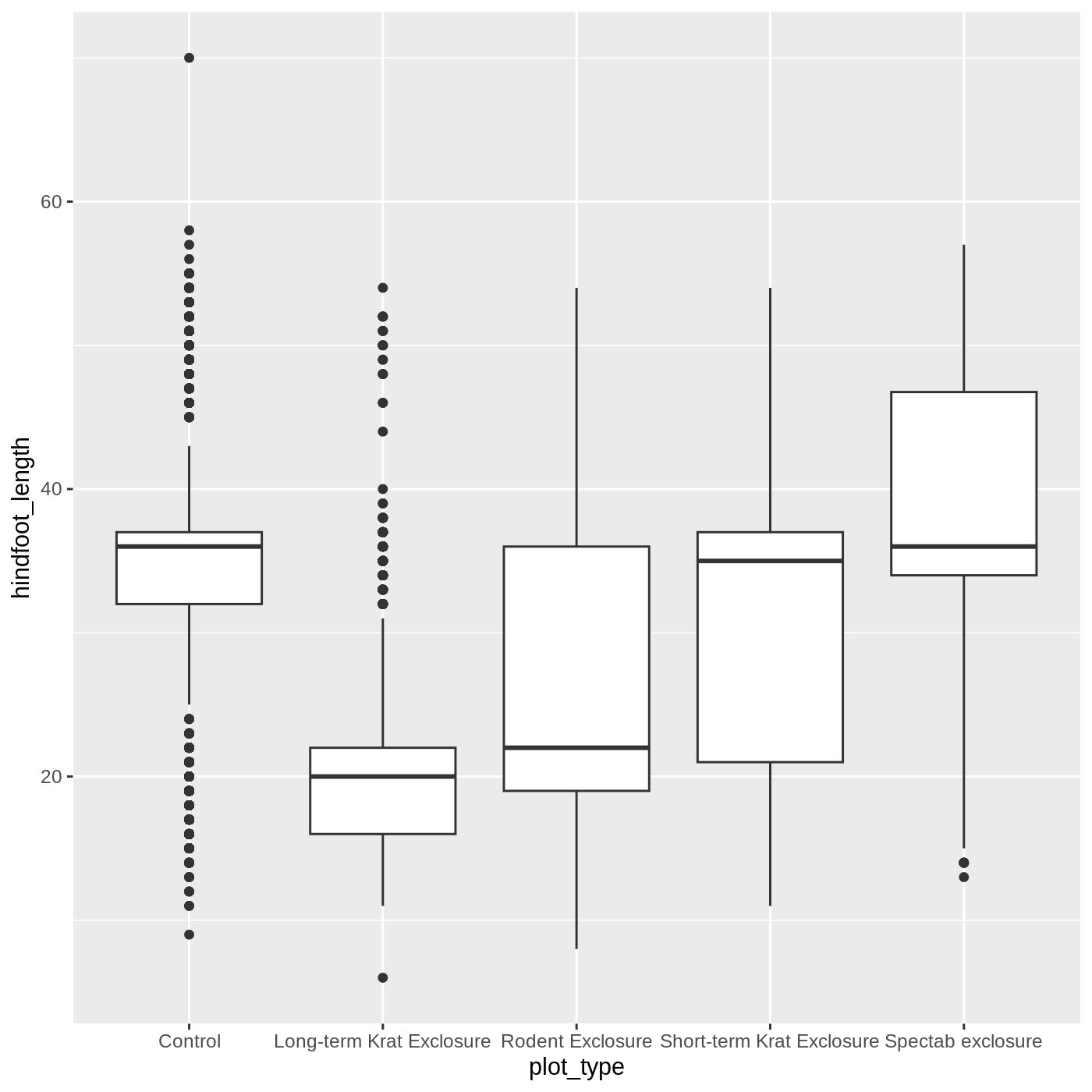
Just as we colored the points before, we can color our boxplot by
plot_type as well:
R
ggplot(data = complete_old, mapping = aes(x = plot_type, y = hindfoot_length, color = plot_type)) +
geom_boxplot()
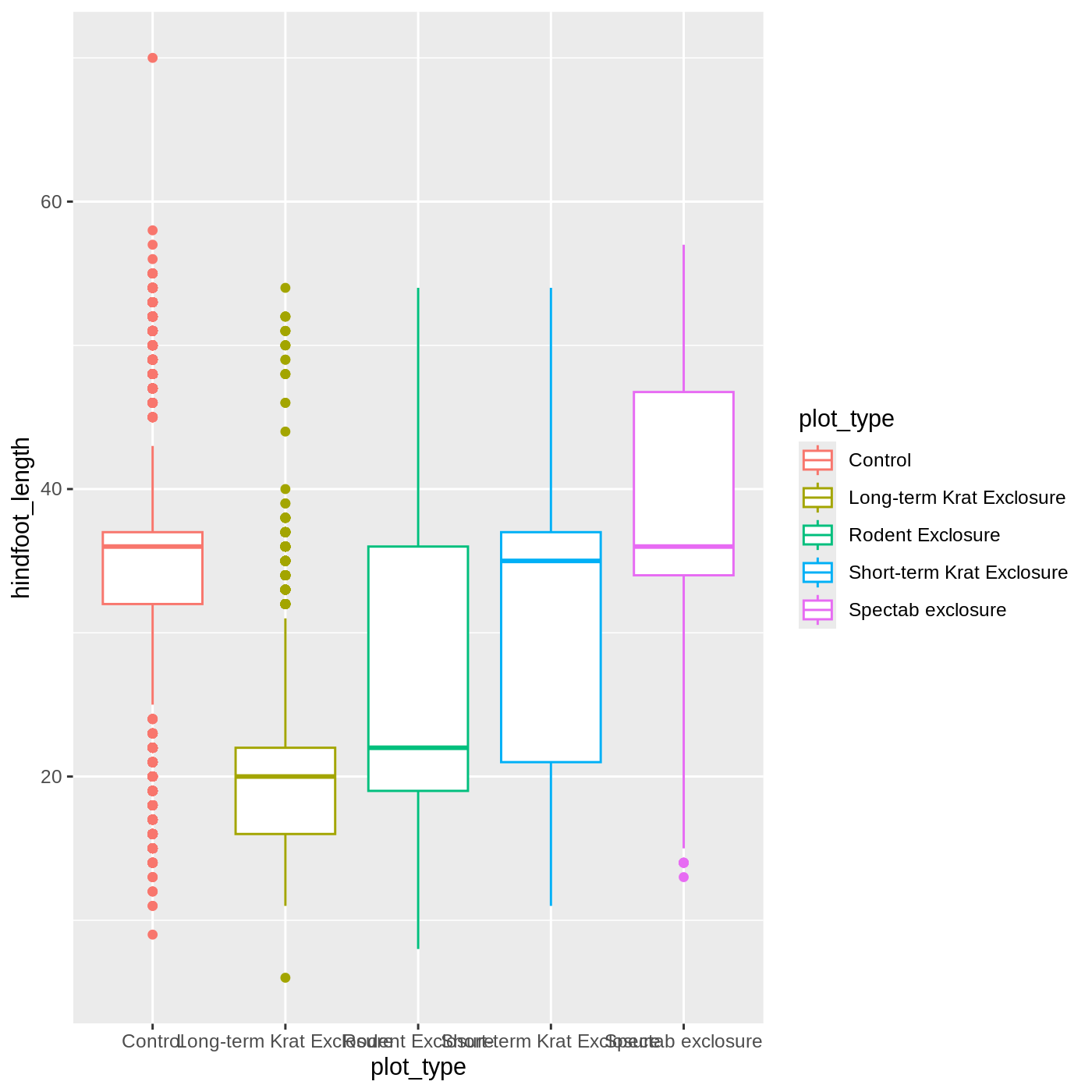
It looks like color has only affected the outlines of
the boxplot, not the rectangular portions. This is because the
color only impacts 1-dimensional parts of a
ggplot: points and lines. To change the color of
2-dimensional parts of a plot, we use fill:
R
ggplot(data = complete_old, mapping = aes(x = plot_type, y = hindfoot_length, fill = plot_type)) +
geom_boxplot()
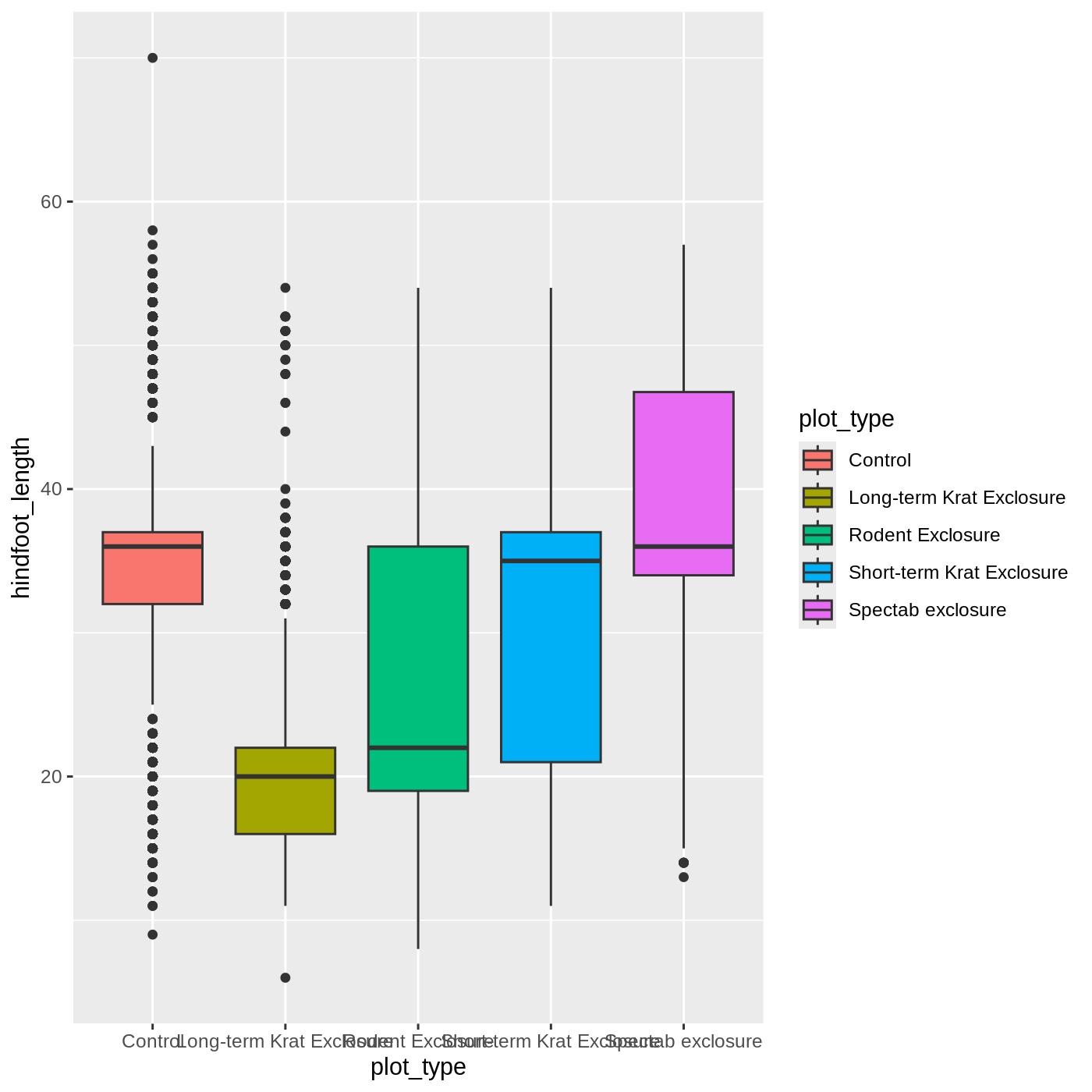
Callout
One thing you may notice is that the axis labels are overlapping each
other, depending on how wide your plot viewer is. One way to help make
them more legible is to wrap the text. We can do that
by modifying the labels for the x axis
scale.
We use the scale_x_discrete() function because we have a
discrete axis, and we modify the labels argument. The
function label_wrap_gen() will wrap the text of the labels
to make them more legible.
R
ggplot(data = complete_old, mapping = aes(x = plot_type, y = hindfoot_length, fill = plot_type)) +
geom_boxplot() +
scale_x_discrete(labels = label_wrap_gen(width = 10))
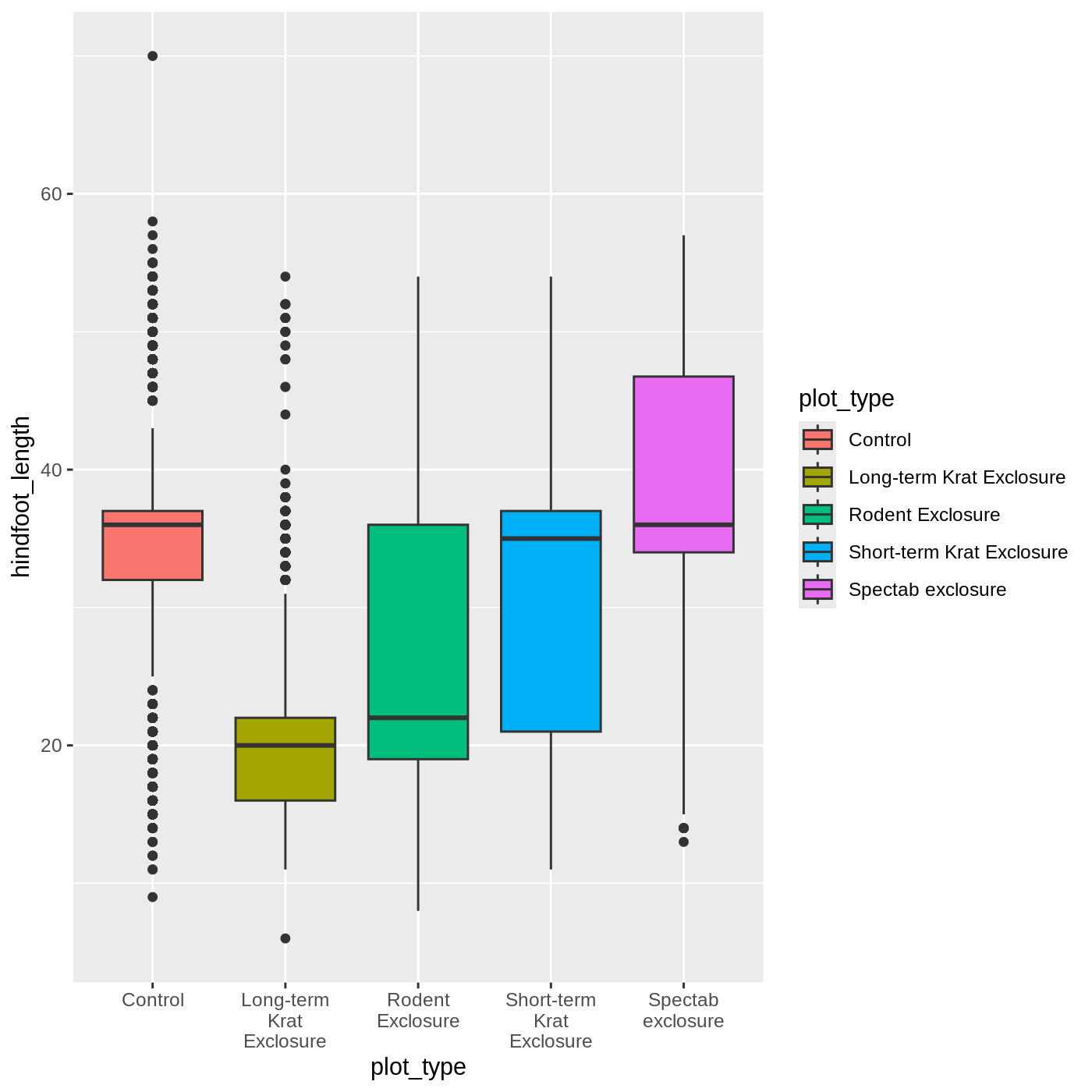
Adding geoms
One of the most powerful aspects of
ggplot is the way we can add components to
a plot in successive layers. While boxplots can be very useful for
summarizing data, it is often helpful to show the raw data as well. With
ggplot, we can easily add another
geom_ to our plot to show the raw data.
Let’s add geom_point() to visualize the raw data. We
will modify the alpha argument to help with
overplotting.
R
ggplot(data = complete_old, mapping = aes(x = plot_type, y = hindfoot_length)) +
geom_boxplot() +
geom_point(alpha = 0.2)
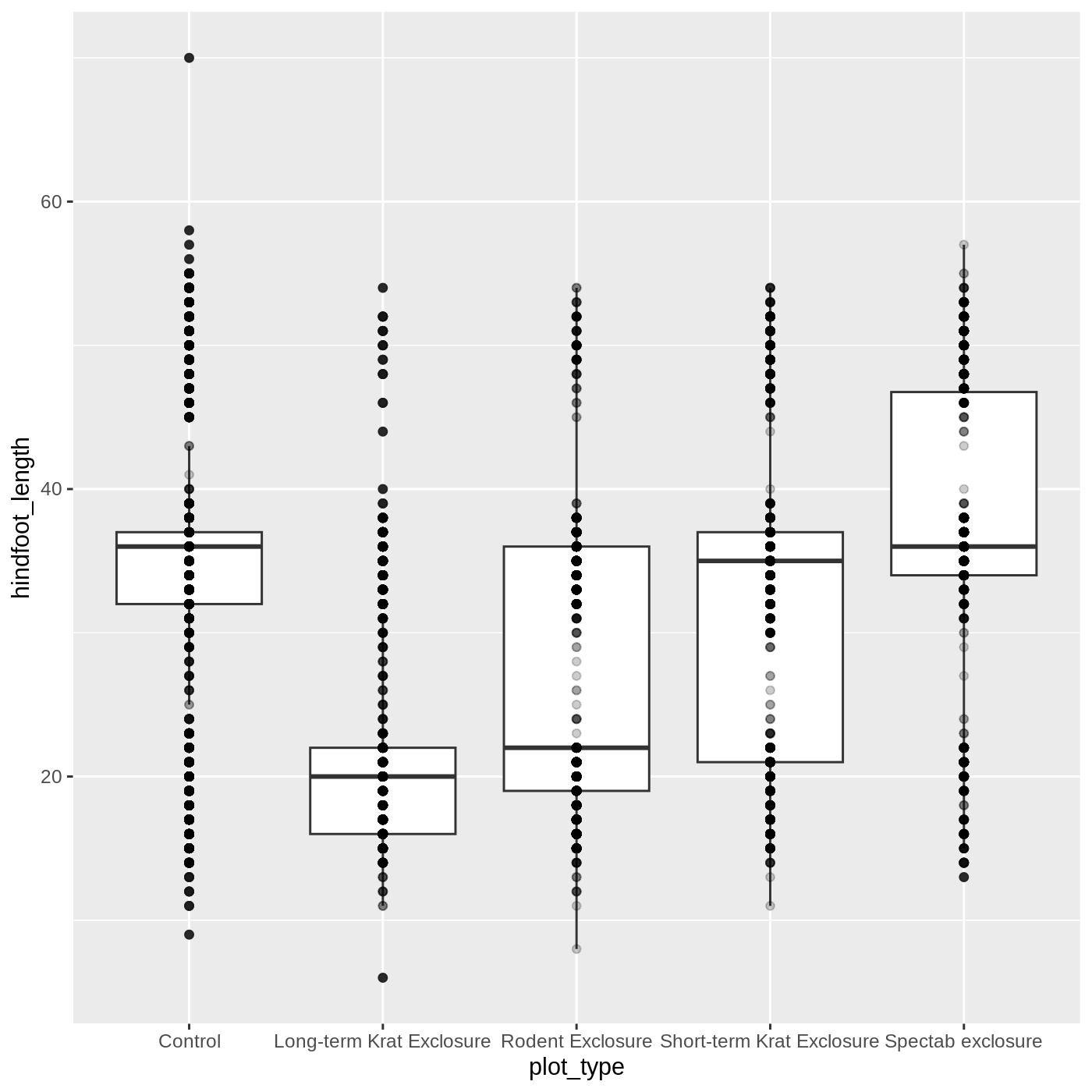
Uh oh… all our points for a given x axis category fall
exactly on a line, which isn’t very useful. We can shift to using
geom_jitter(), which will add points with a bit of random
noise added to the positions to prevent this from happening.
R
ggplot(data = complete_old, mapping = aes(x = plot_type, y = hindfoot_length)) +
geom_boxplot() +
geom_jitter(alpha = 0.2)
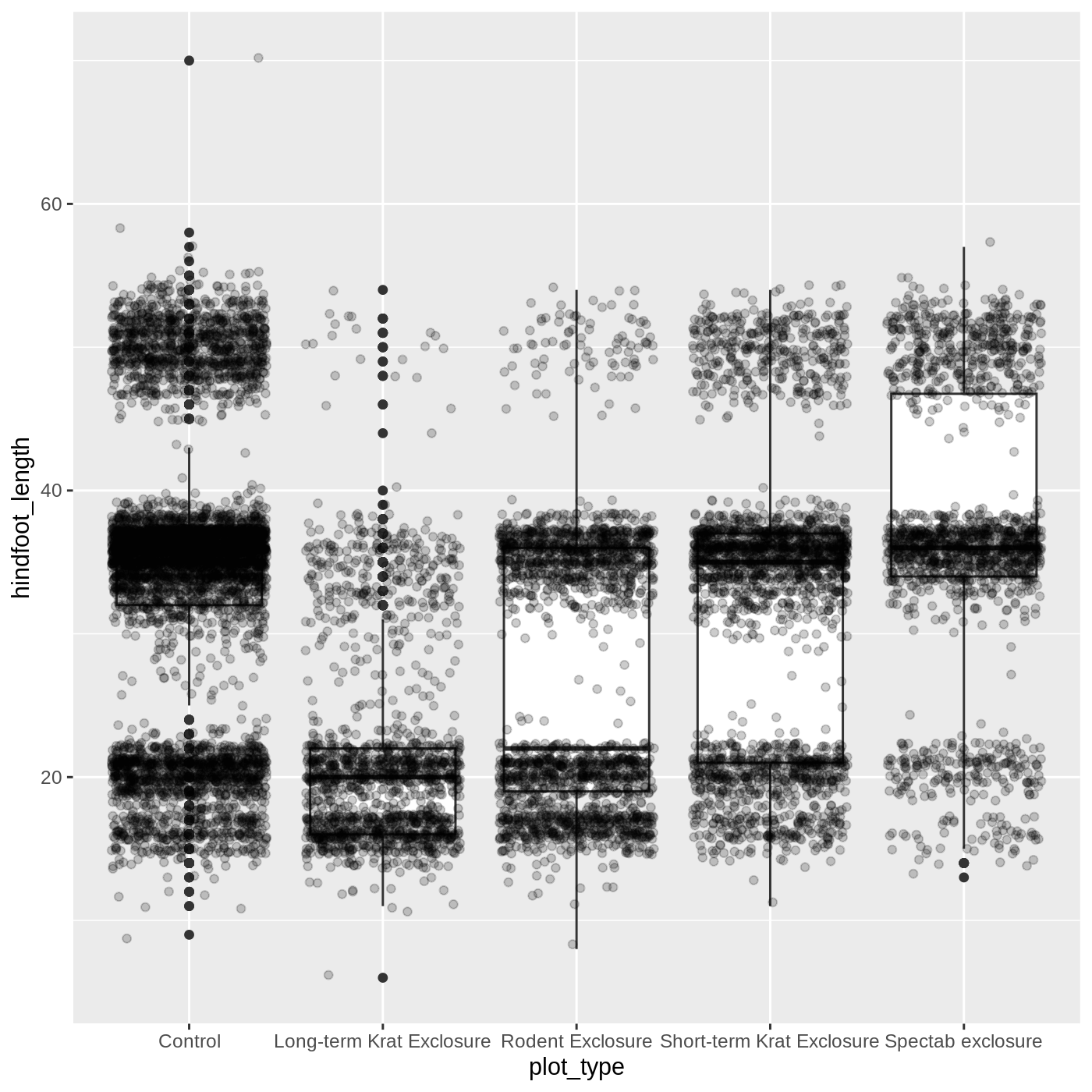
You may have noticed that some of our data points are now appearing
on our plot twice: the outliers are plotted as black points from
geom_boxplot(), but they are also plotted with
geom_jitter(). Since we don’t want to represent these data
multiple times in the same form (points), we can stop
geom_boxplot() from plotting them. We do this by setting
the outlier.shape argument to NA, which means
the outliers don’t have a shape to be plotted.
R
ggplot(data = complete_old, mapping = aes(x = plot_type, y = hindfoot_length)) +
geom_boxplot(outlier.shape = NA) +
geom_jitter(alpha = 0.2)
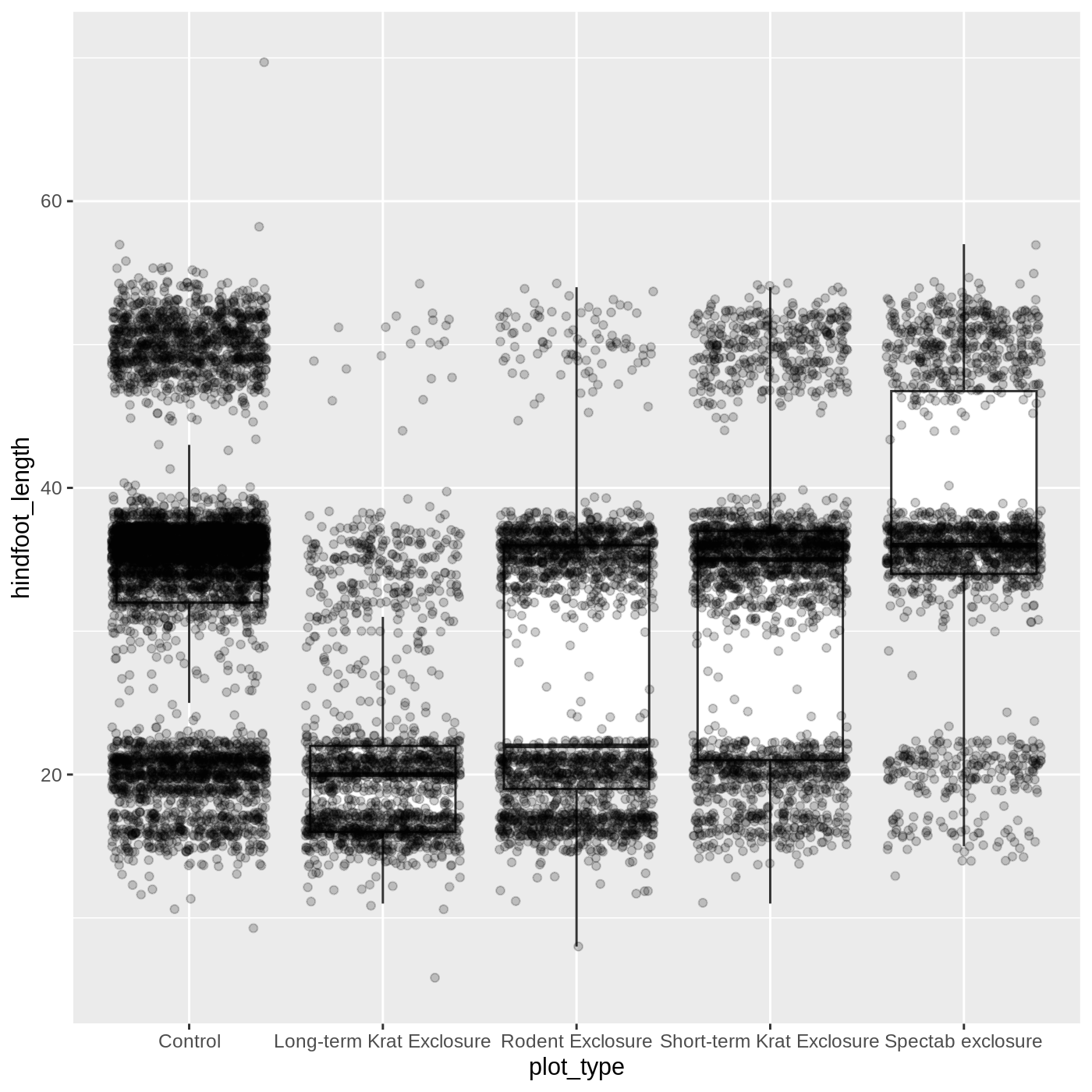
Just as before, we can map plot_type to
color by putting it inside aes().
R
ggplot(data = complete_old, mapping = aes(x = plot_type, y = hindfoot_length, color = plot_type)) +
geom_boxplot(outlier.shape = NA) +
geom_jitter(alpha = 0.2)
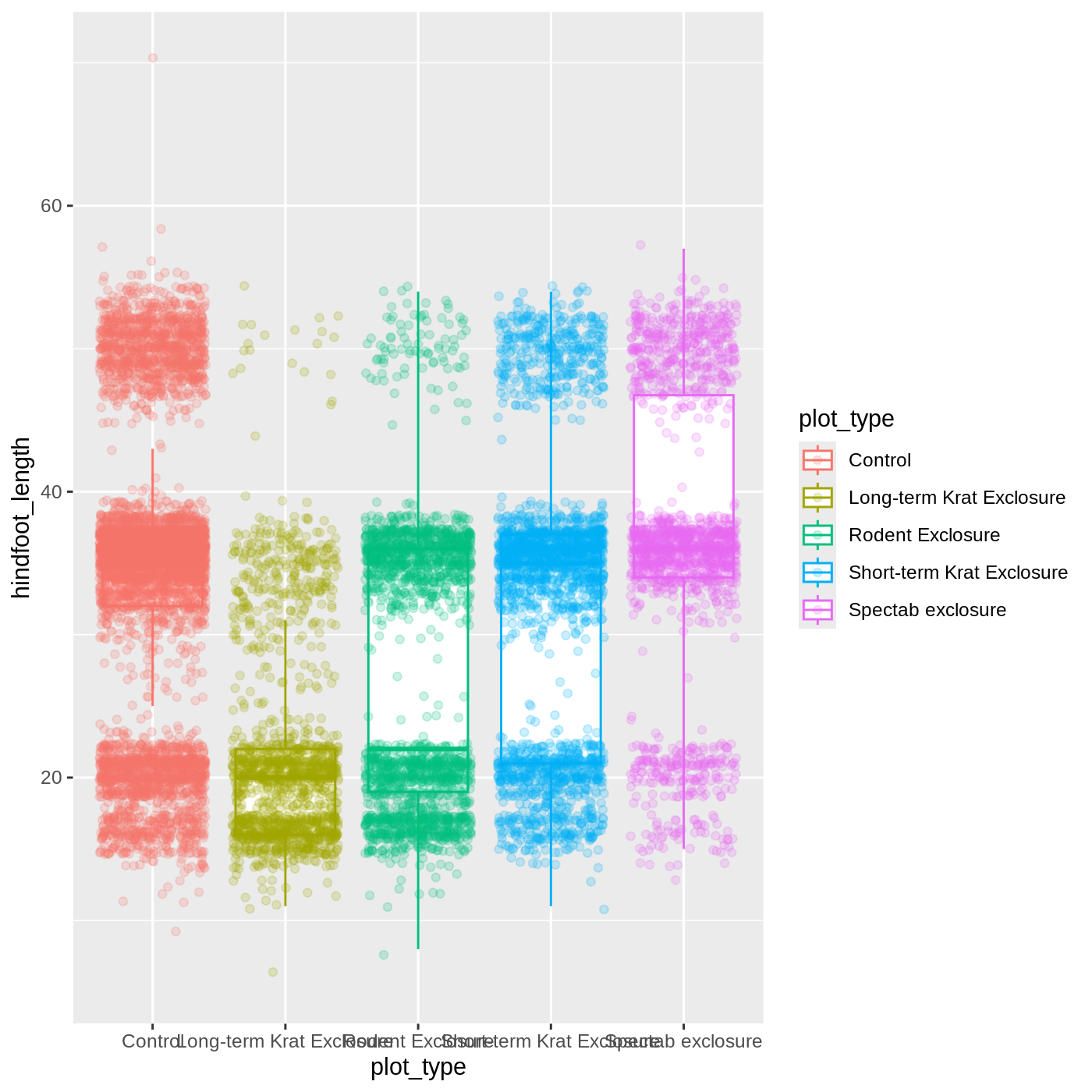
Notice that both the color of the points and the color of the boxplot
lines changed. Any time we specify an aes() mapping inside
our initial ggplot() function, that mapping will apply to
all our geoms.
If we want to limit the mapping to a single geom, we can
put the mapping into the specific geom_ function, like
this:
R
ggplot(data = complete_old, mapping = aes(x = plot_type, y = hindfoot_length)) +
geom_boxplot(outlier.shape = NA) +
geom_jitter(aes(color = plot_type), alpha = 0.2)
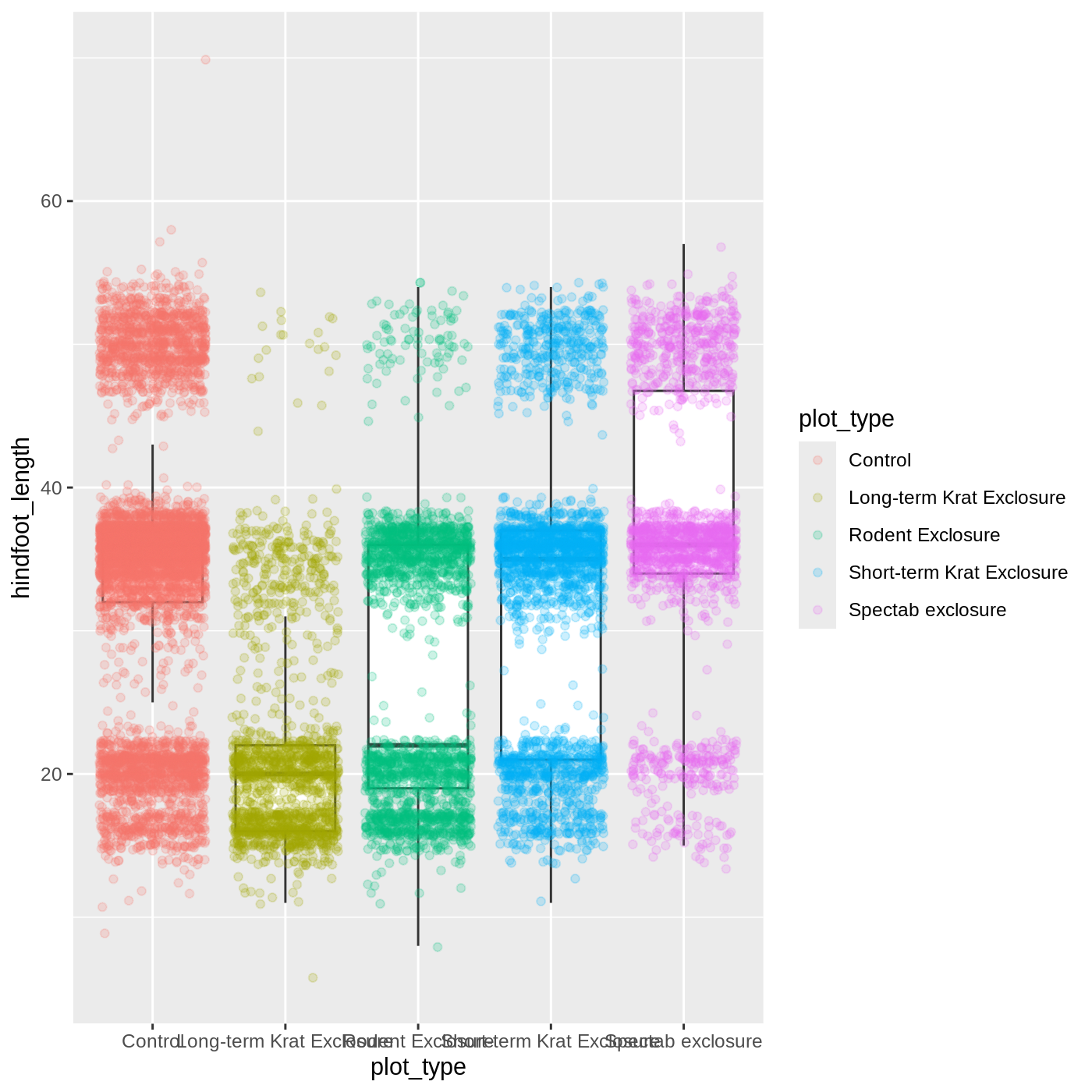
Now our points are colored according to plot_type, but
the boxplots are all the same color. One thing you might notice is that
even with alpha = 0.2, the points obscure parts of the
boxplot. This is because the geom_point() layer comes after
the geom_boxplot() layer, which means the points are
plotted on top of the boxes. To put the boxplots on top, we switch the
order of the layers:
R
ggplot(data = complete_old, mapping = aes(x = plot_type, y = hindfoot_length)) +
geom_jitter(aes(color = plot_type), alpha = 0.2) +
geom_boxplot(outlier.shape = NA)
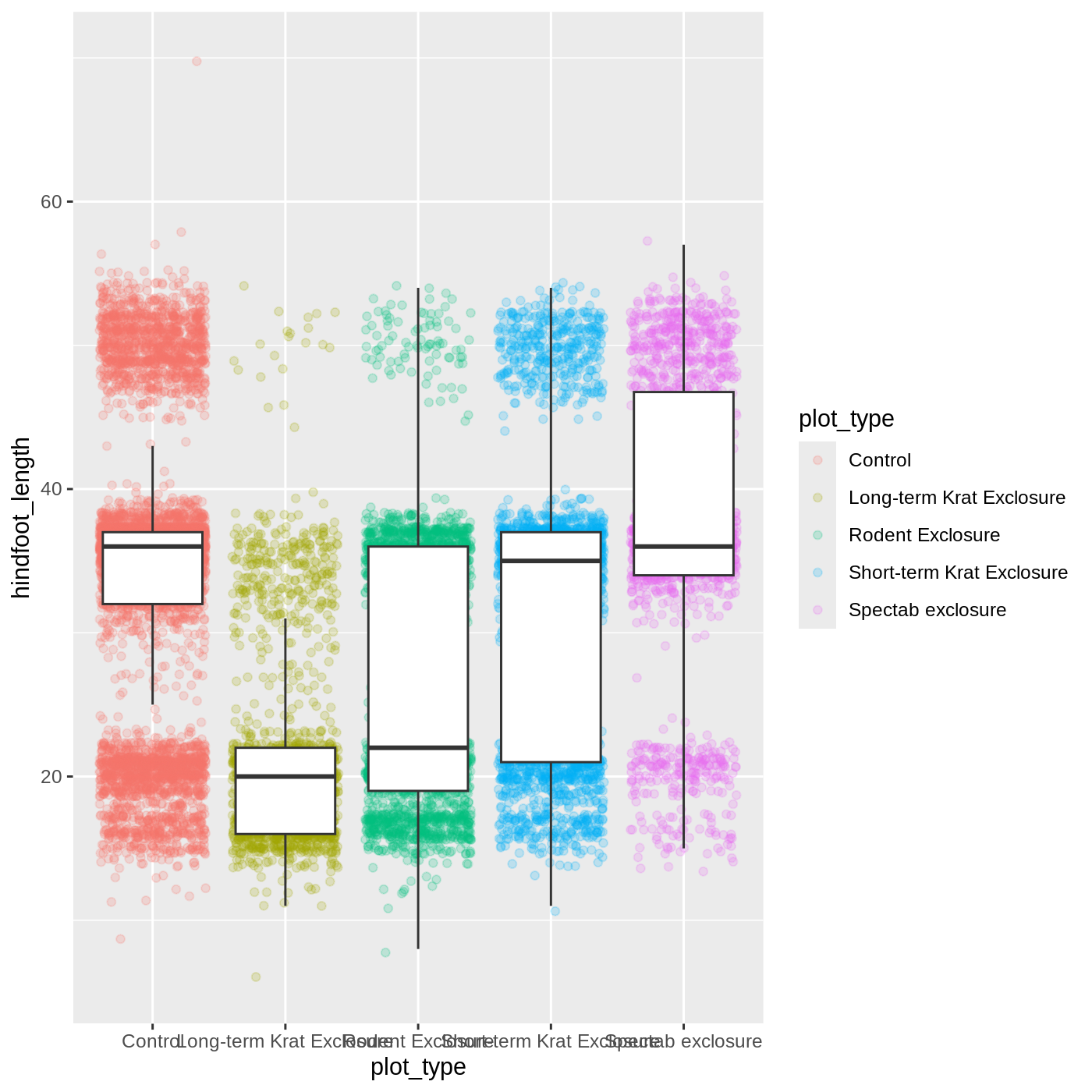
Now we have the opposite problem! The white fill of the
boxplots completely obscures some of the points. To address this
problem, we can remove the fill from the boxplots
altogether, leaving only the black lines. To do this, we set
fill to NA:
R
ggplot(data = complete_old, mapping = aes(x = plot_type, y = hindfoot_length)) +
geom_jitter(aes(color = plot_type), alpha = 0.2) +
geom_boxplot(outlier.shape = NA, fill = NA)
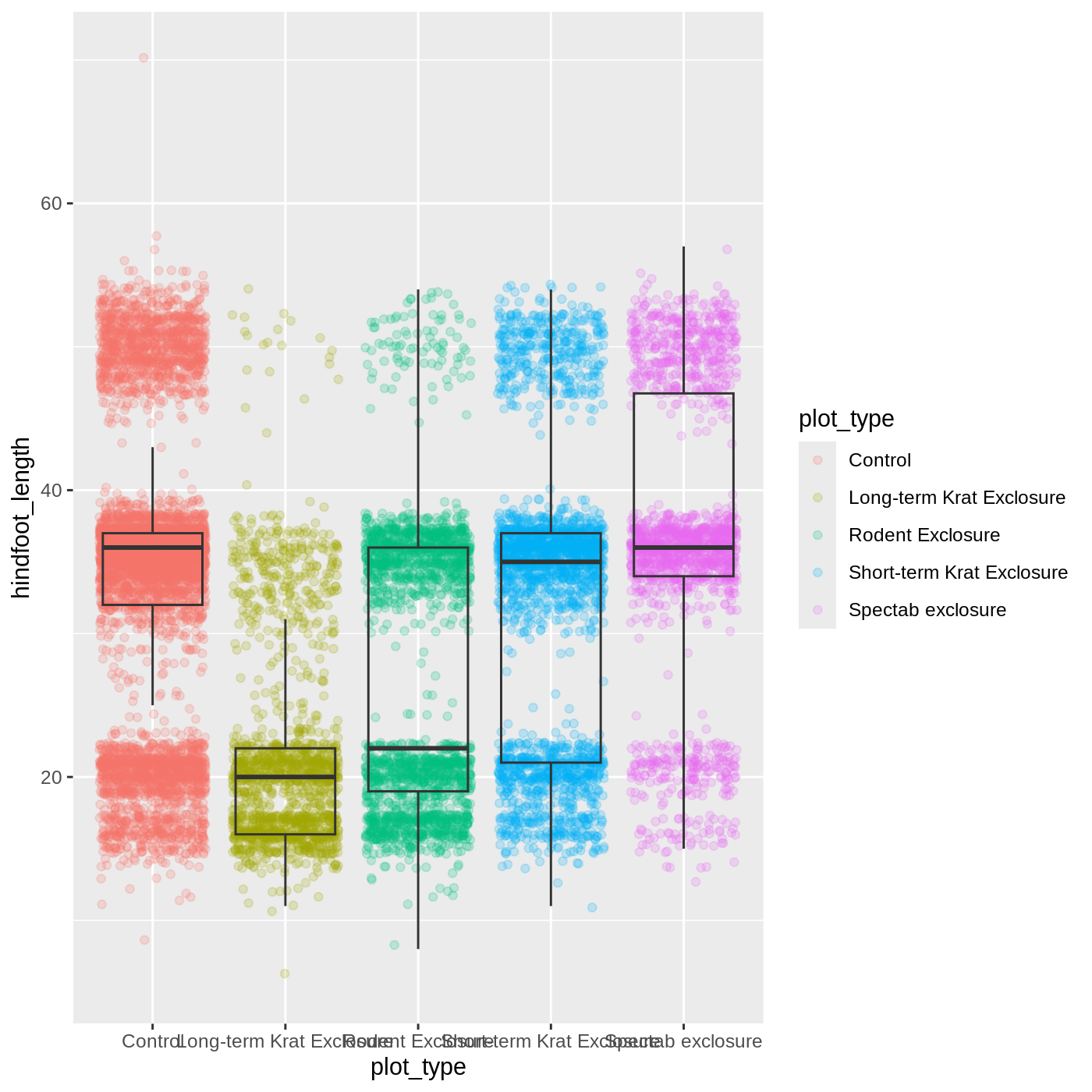
Now we can see all the raw data and our boxplots on top.
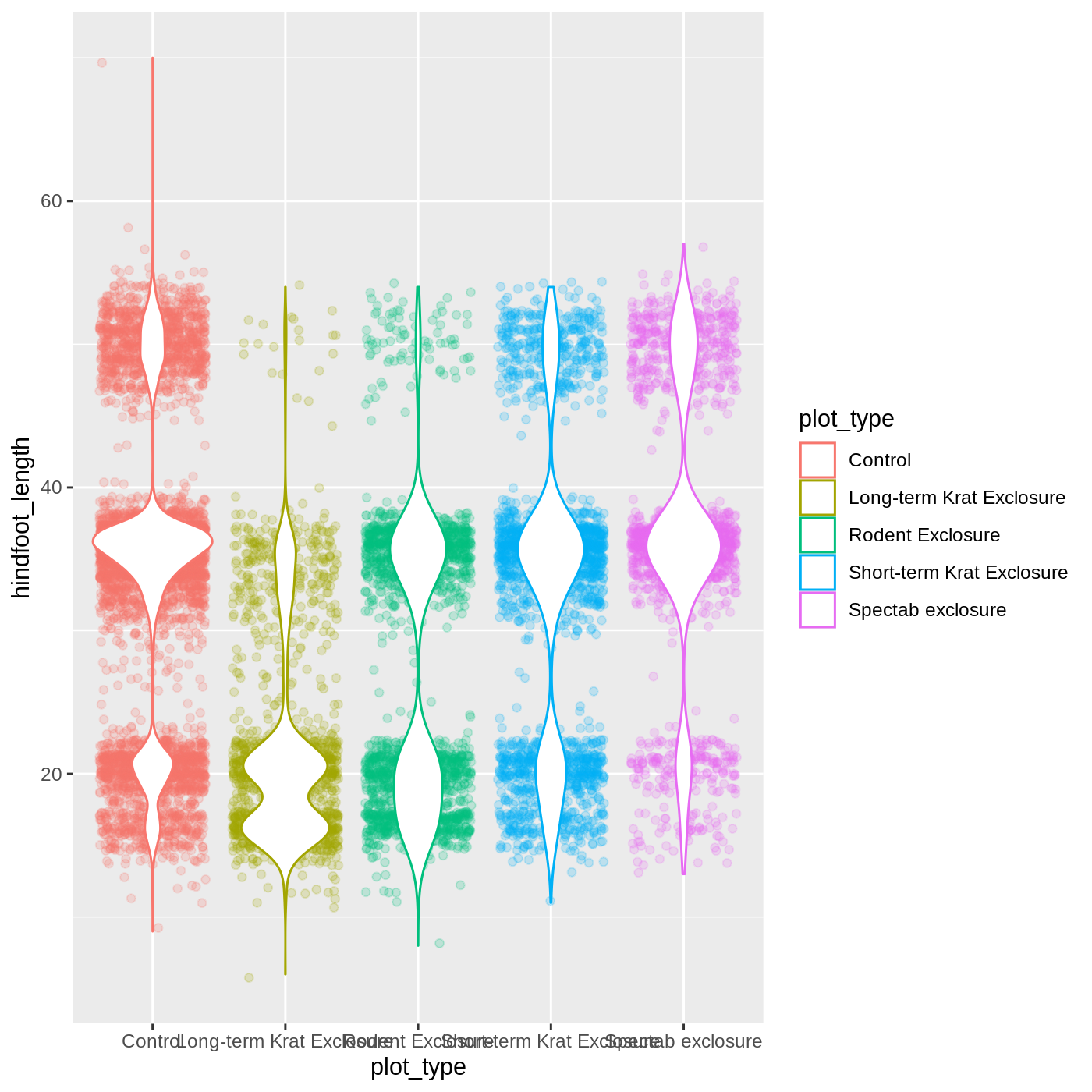
Challenge 2: Change geoms
Violin plots are similar to boxplots- try making one using
plot_type and hindfoot_length as the x and y
variables. Remember that all geom functions start with
geom_, followed by the type of geom.
This might also be a place to test your search engine skills. It is
often useful to search for
R package_name stuff you want to search. So for this
example we might search for R ggplot2 violin plot.
R
ggplot(data = complete_old,
mapping = aes(x = plot_type,
y = hindfoot_length,
color = plot_type)) +
geom_jitter(alpha = 0.2) +
geom_violin(fill = "white")
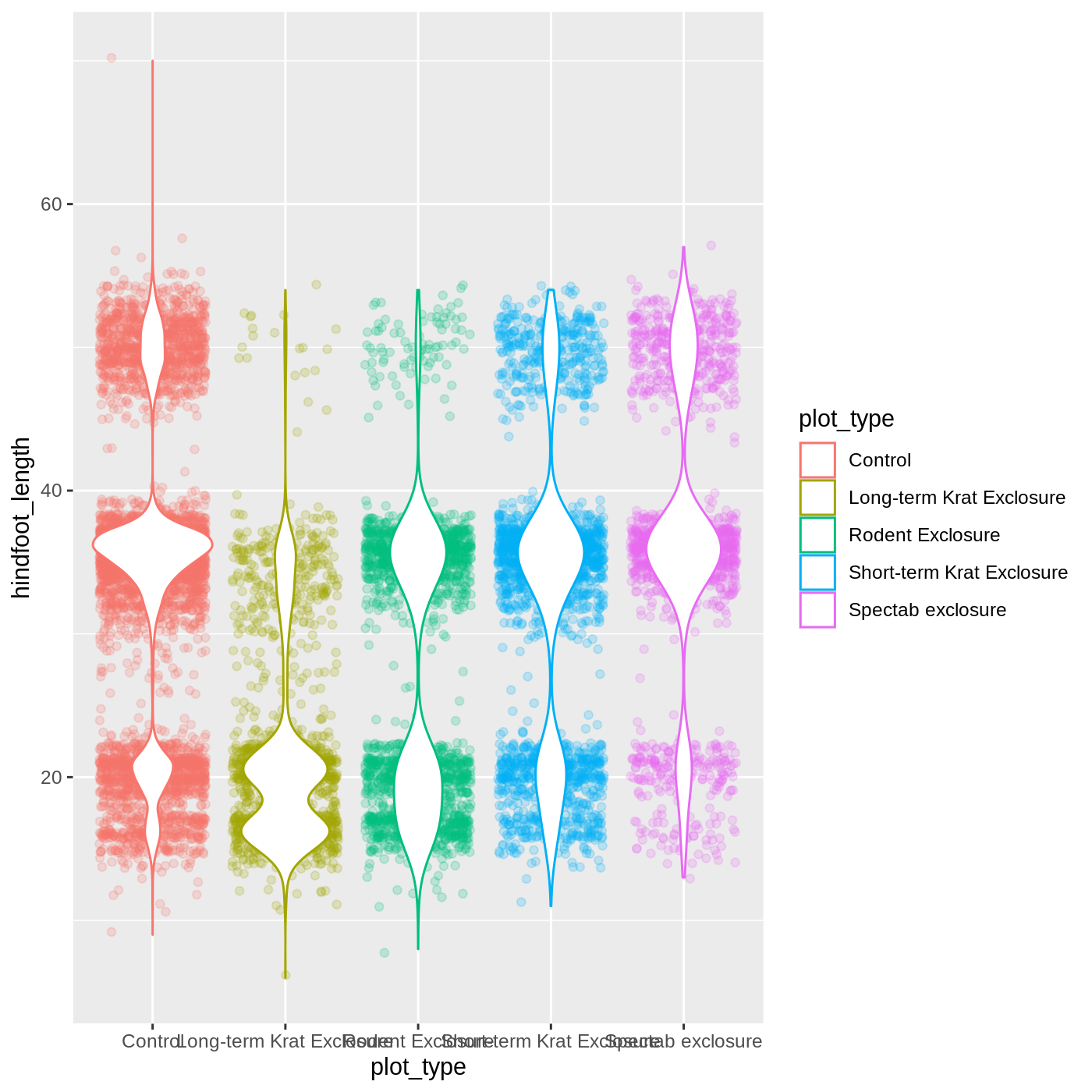
R
ggplot(data = complete_old,
mapping = aes(x = plot_type,
y = hindfoot_length,
color = plot_type)) +
geom_jitter(alpha = 0.2) +
geom_violin(fill = "white")
Changing themes
So far we’ve been changing the appearance of parts of our plot
related to our data and the geom_ functions, but we can
also change many of the non-data components of our plot.
At this point, we are pretty happy with the basic layout of our plot,
so we can assign it to a plot to a named
object. We do this using the assignment
arrow <-. What we are doing here is taking the
result of the code on the right side of the arrow, and assigning it to
an object whose name is on the left side of the arrow.
We will create an object called myplot. If you run the
name of the ggplot2 object, it will show the plot, just
like if you ran the code itself.
R
myplot <- ggplot(data = complete_old, mapping = aes(x = plot_type, y = hindfoot_length)) +
geom_jitter(aes(color = plot_type), alpha = 0.2) +
geom_boxplot(outlier.shape = NA, fill = NA)
myplot
WARNING
Warning: Removed 2733 rows containing non-finite outside the scale range
(`stat_boxplot()`).WARNING
Warning: Removed 2733 rows containing missing values or values outside the scale range
(`geom_point()`).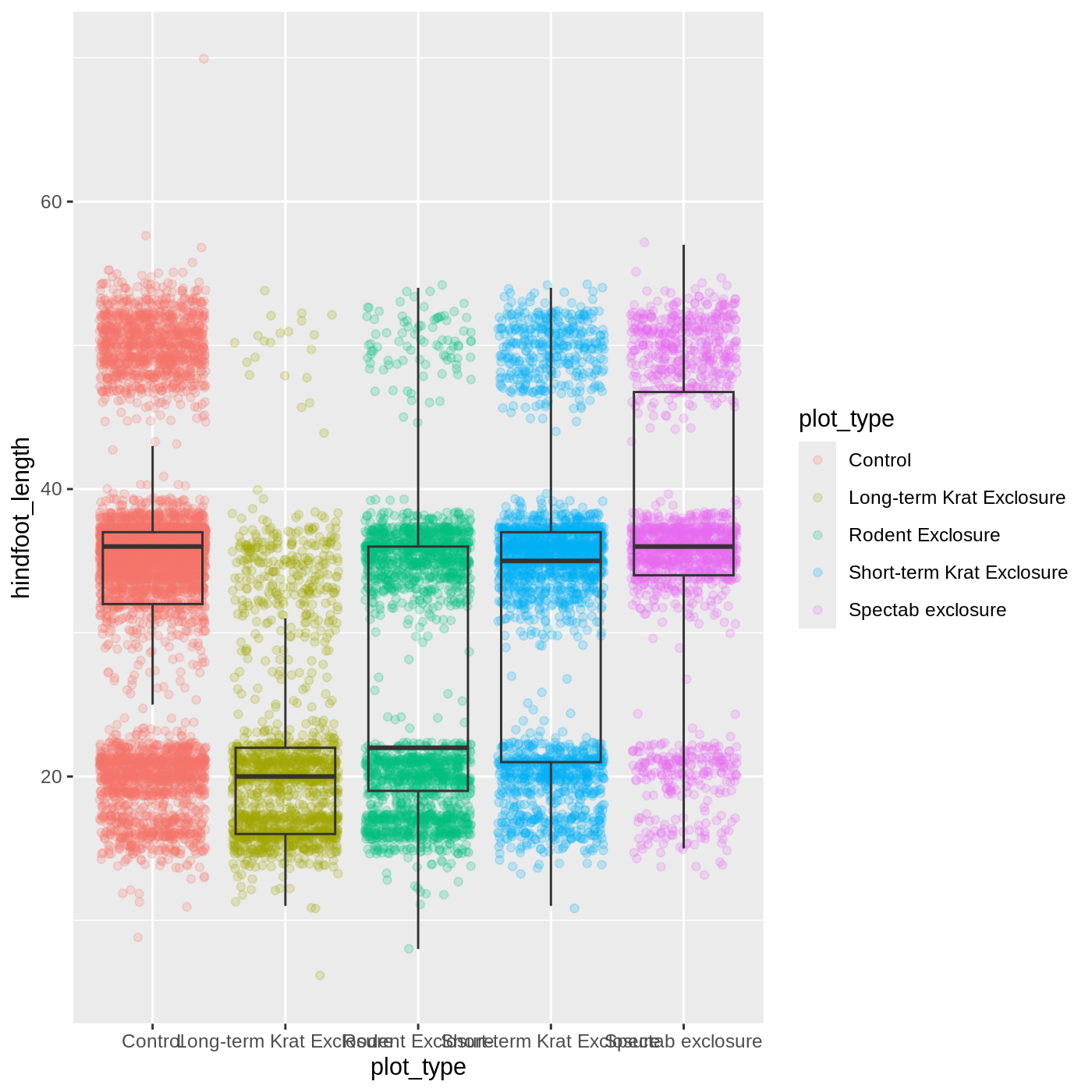
This process of assigning something to an object is
not specific to ggplot2, but rather a general feature of R.
We will be using it a lot in the rest of this lesson. We can now work
with the myplot object as if it was a block of
ggplot2 code, which means we can use + to add
new components to it.
We can change the overall appearance using theme_
functions. Let’s try a black-and-white theme by adding
theme_bw() to our plot:
R
myplot + theme_bw()
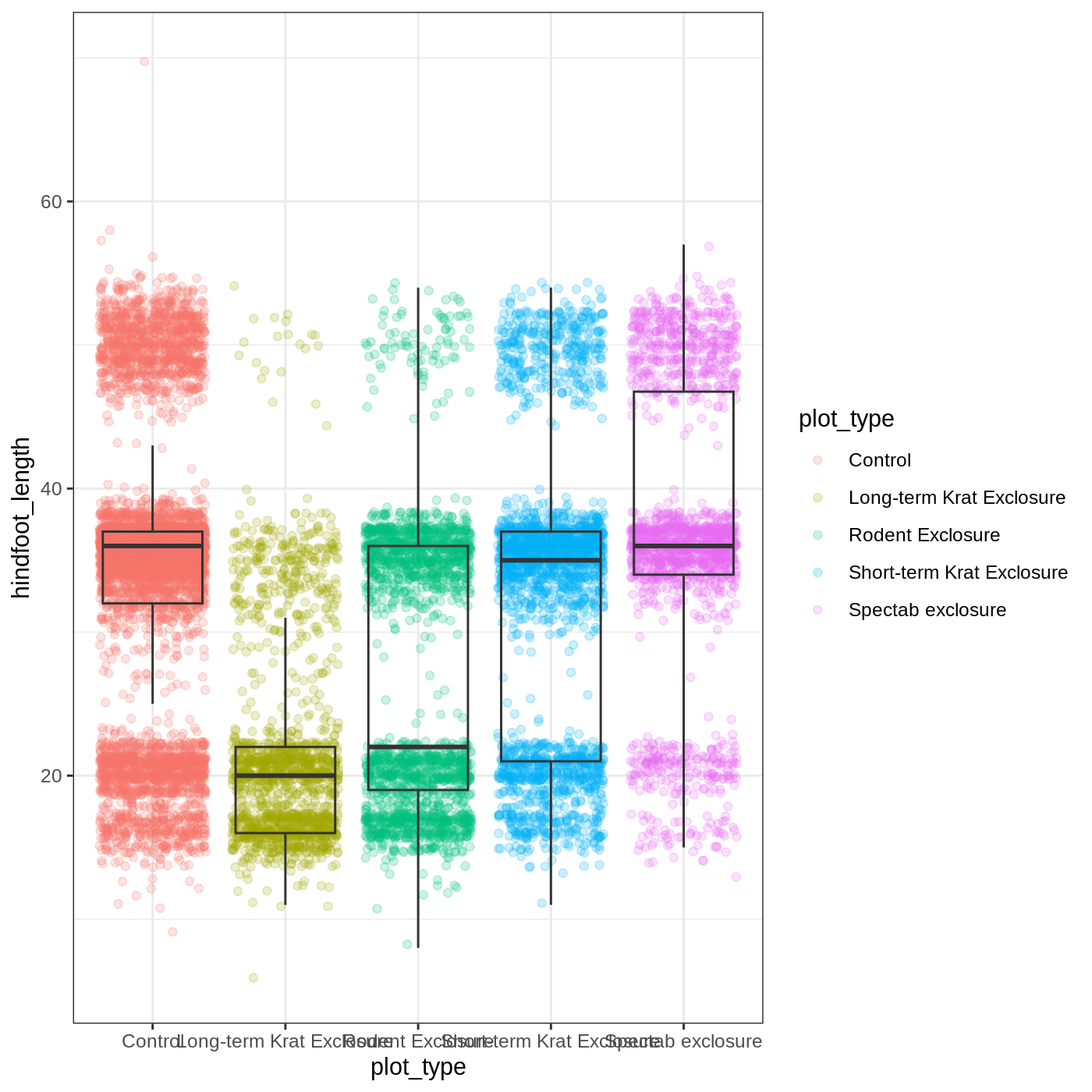
As you can see, a number of parts of the plot have changed.
theme_ functions usually control many aspects of a plot’s
appearance all at once, for the sake of convenience. To individually
change parts of a plot, we can use the theme() function,
which can take many different arguments to change things about the text,
grid lines, background color, and more. Let’s try changing the size of
the text on our axis titles. We can do this by specifying that the
axis.title should be an element_text() with
size set to 14.
R
myplot +
theme_bw() +
theme(axis.title = element_text(size = 14))
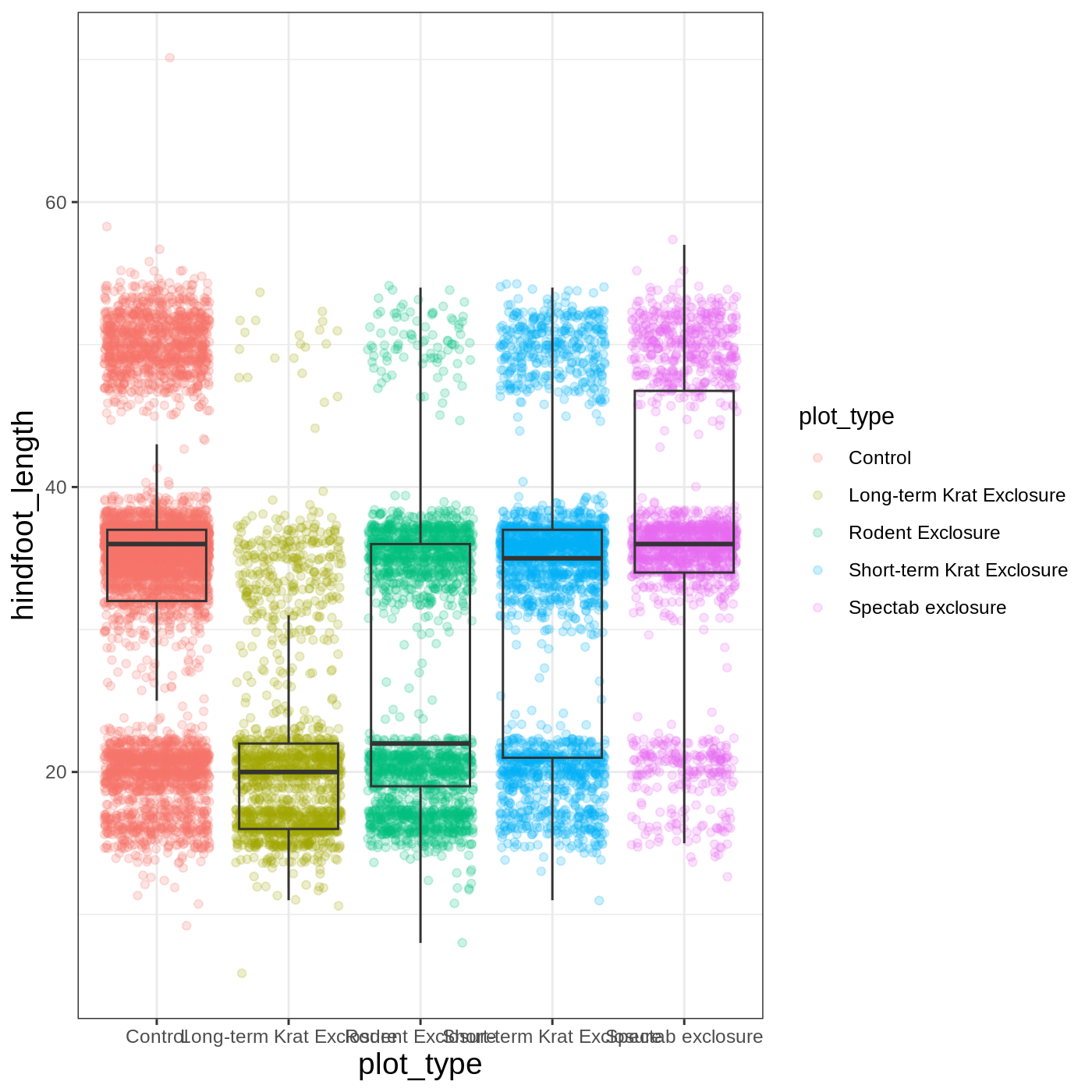
Another change we might want to make is to remove the vertical grid
lines. Since our x axis is categorical, those grid lines aren’t useful.
To do this, inside theme(), we will change the
panel.grid.major.x to an element_blank().
R
myplot +
theme_bw() +
theme(axis.title = element_text(size = 14),
panel.grid.major.x = element_blank())
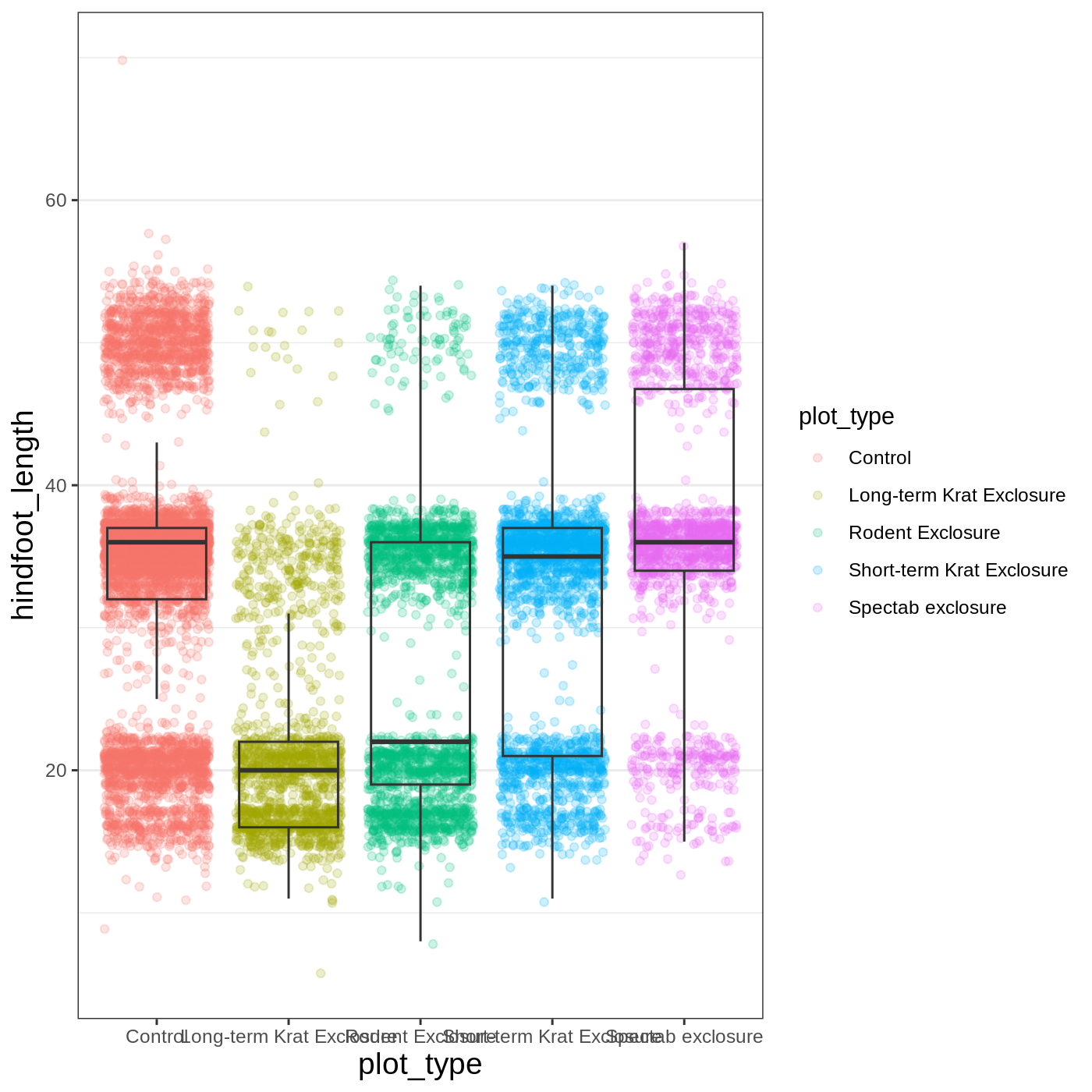
Another useful change might be to remove the color legend, since that
information is already on our x axis. For this one, we will set
legend.position to “none”.
R
myplot +
theme_bw() +
theme(axis.title = element_text(size = 14),
panel.grid.major.x = element_blank(),
legend.position = "none")
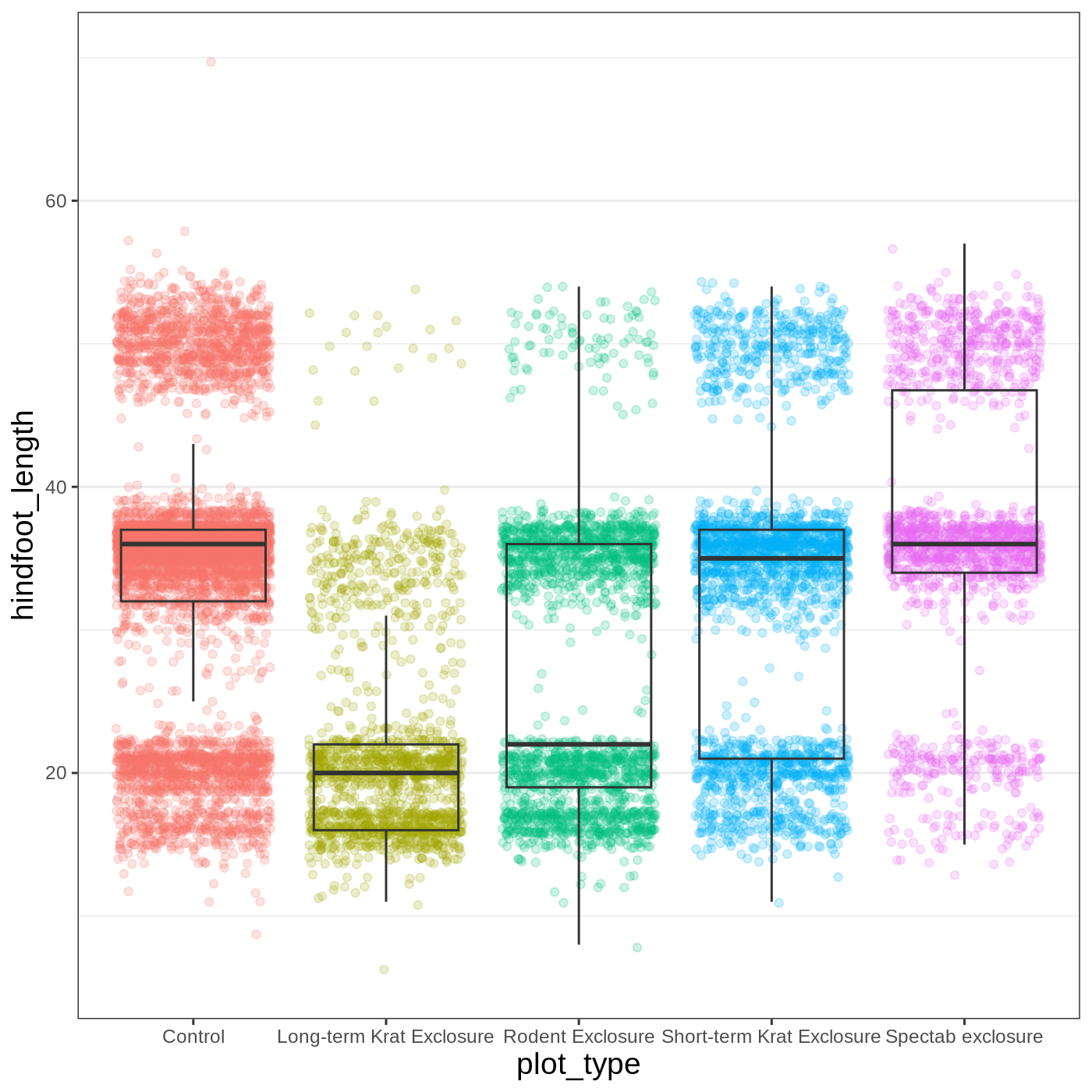
Callout
Because there are so many possible arguments to the
theme() function, it can sometimes be hard to find the
right one. Here are some tips for figuring out how to modify a plot
element:
- type out
theme(), put your cursor between the parentheses, and hit Tab to bring up a list of arguments- you can scroll through the arguments, or start typing, which will shorten the list of potential matches
- like many things in the
tidyverse, similar argument start with similar names- there are
axis,legend,panel,plot, andstriparguments
- there are
- arguments have hierarchy
-
textcontrols all text in the whole plot -
axis.titlecontrols the text for the axis titles -
axis.title.xcontrols the text for the x axis title
-
Callout
You may have noticed that we have used 3 different approaches to
getting rid of something in ggplot:
-
outlier.shape = NAto remove the outliers from our boxplot -
panel.grid.major.x = element_blank()to remove the x grid lines -
legend.position = "none"to remove our legend
Why are there so many ways to do what seems like the same thing?? This is a common frustration when working with R, or with any programming language. There are a couple reasons for it:
- Different people contribute to different packages and functions, and they may choose to do things differently.
- Code may appear to be doing the same thing, when the
details are actually quite different. The inner workings of
ggplot2are actually quite complex, since it turns out making plots is a very complicated process! Because of this, things that seem the same (removing parts of a plot), may actually be operating on very different components or stages of the final plot. - Developing packages is a highly iterative process, and sometimes
things change. However, changing too much stuff can make old code break.
Let’s say removing the legend was introduced as a feature of
ggplot2, and then a lot of time passed before someone added the feature letting you remove outliers fromgeom_boxplot(). Changing the way you remove the legend, so that it’s the same as the boxplot approach, could break all of the code written in the meantime, so developers may opt to keep the old approach in place.
Changing labels
Our plot is really shaping up now. However, we probably want to make
our axis titles nicer, and perhaps add a main title to the plot. We can
do this using the labs() function:
R
myplot +
theme_bw() +
theme(axis.title = element_text(size = 14),
legend.position = "none") +
labs(title = "Rodent size by plot type",
x = "Plot type",
y = "Hindfoot length (mm)")
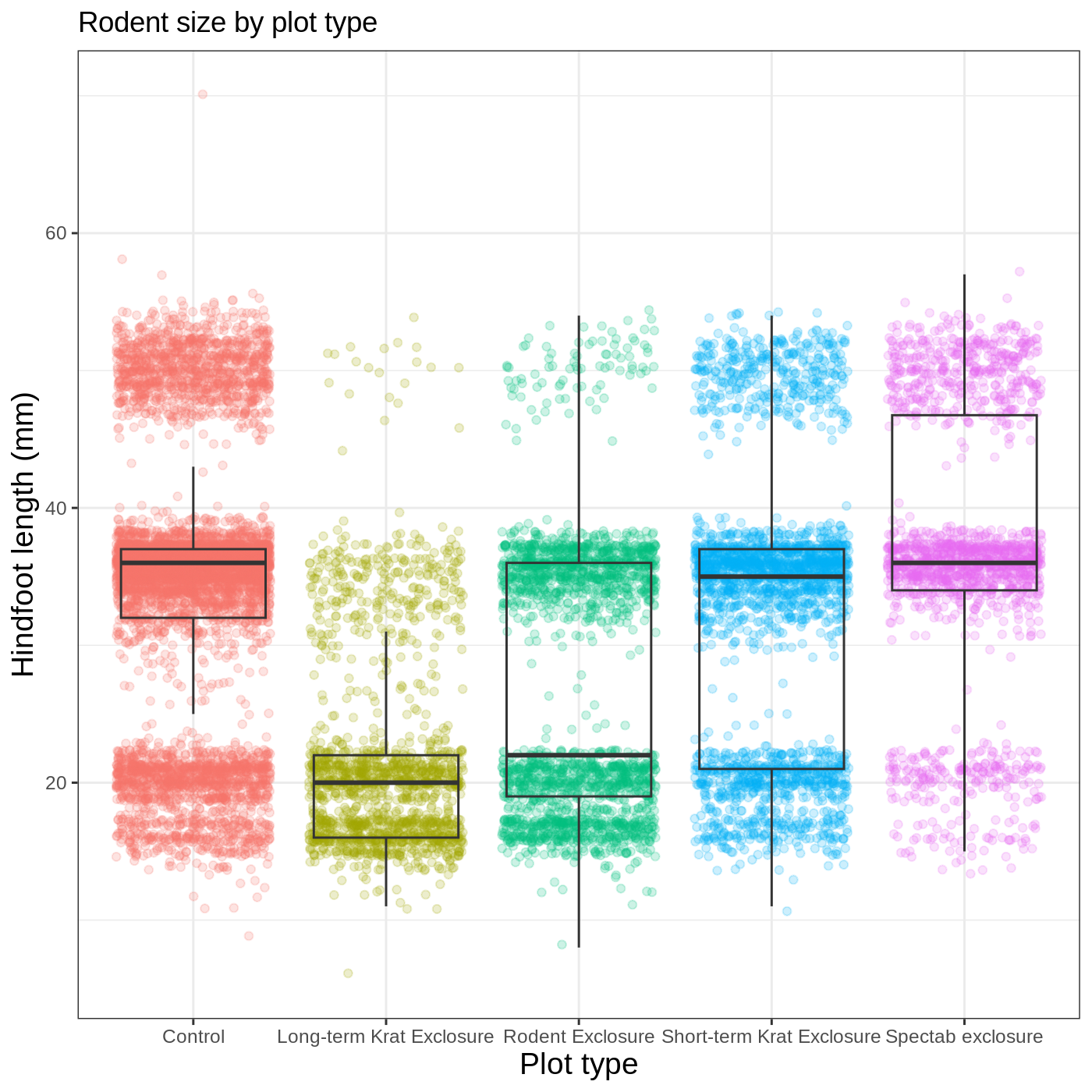
We removed our legend from this plot, but you can also change the
titles of various legends using labs(). For example,
labs(color = "Plot type") would change the title of a color
scale legend to “Plot type”.
R
myplot +
theme_bw() +
theme(axis.title = element_text(size = 14), legend.position = "none",
plot.title = element_text(face = "bold", size = 20)) +
labs(title = "Rodent size by plot type",
subtitle = "Long-term dataset from Portal, AZ",
x = "Plot type",
y = "Hindfoot length (mm)")
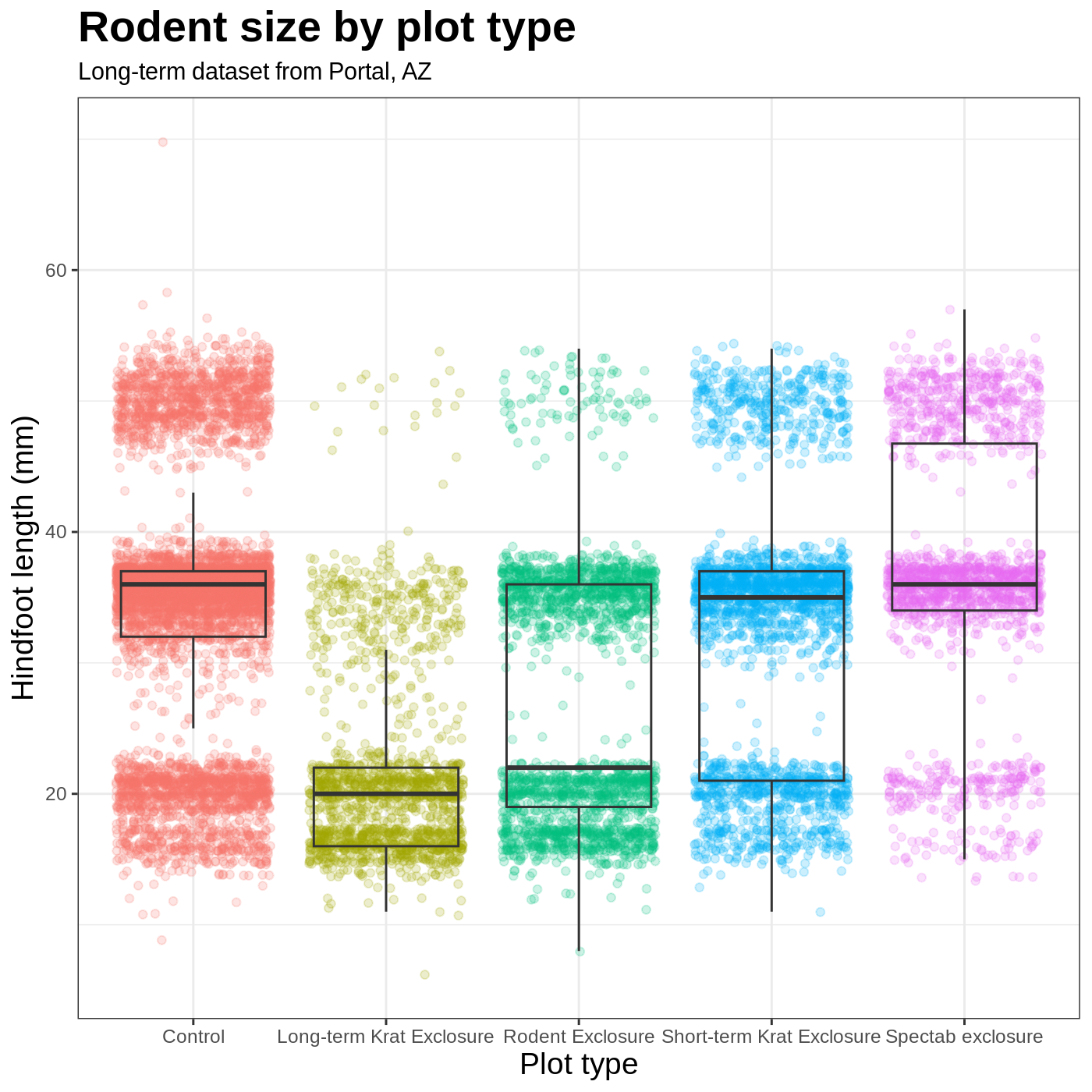
Faceting
One of the most powerful features of
ggplot is the ability to quickly split a
plot into multiple smaller plots based on a categorical variable, which
is called faceting.
So far we’ve mapped variables to the x axis, the y axis, and color, but trying to add a 4th variable becomes difficult. Changing the shape of a point might work, but only for very few categories, and even then, it can be hard to tell the differences between the shapes of small points.
Instead of cramming one more variable into a single plot, we will use
the facet_wrap() function to generate a series of smaller
plots, split out by sex. We also use ncol to
specify that we want them arranged in a single column:
R
myplot +
theme_bw() +
theme(axis.title = element_text(size = 14),
legend.position = "none",
panel.grid.major.x = element_blank()) +
labs(title = "Rodent size by plot type",
x = "Plot type",
y = "Hindfoot length (mm)",
color = "Plot type") +
facet_wrap(vars(sex), ncol = 1)
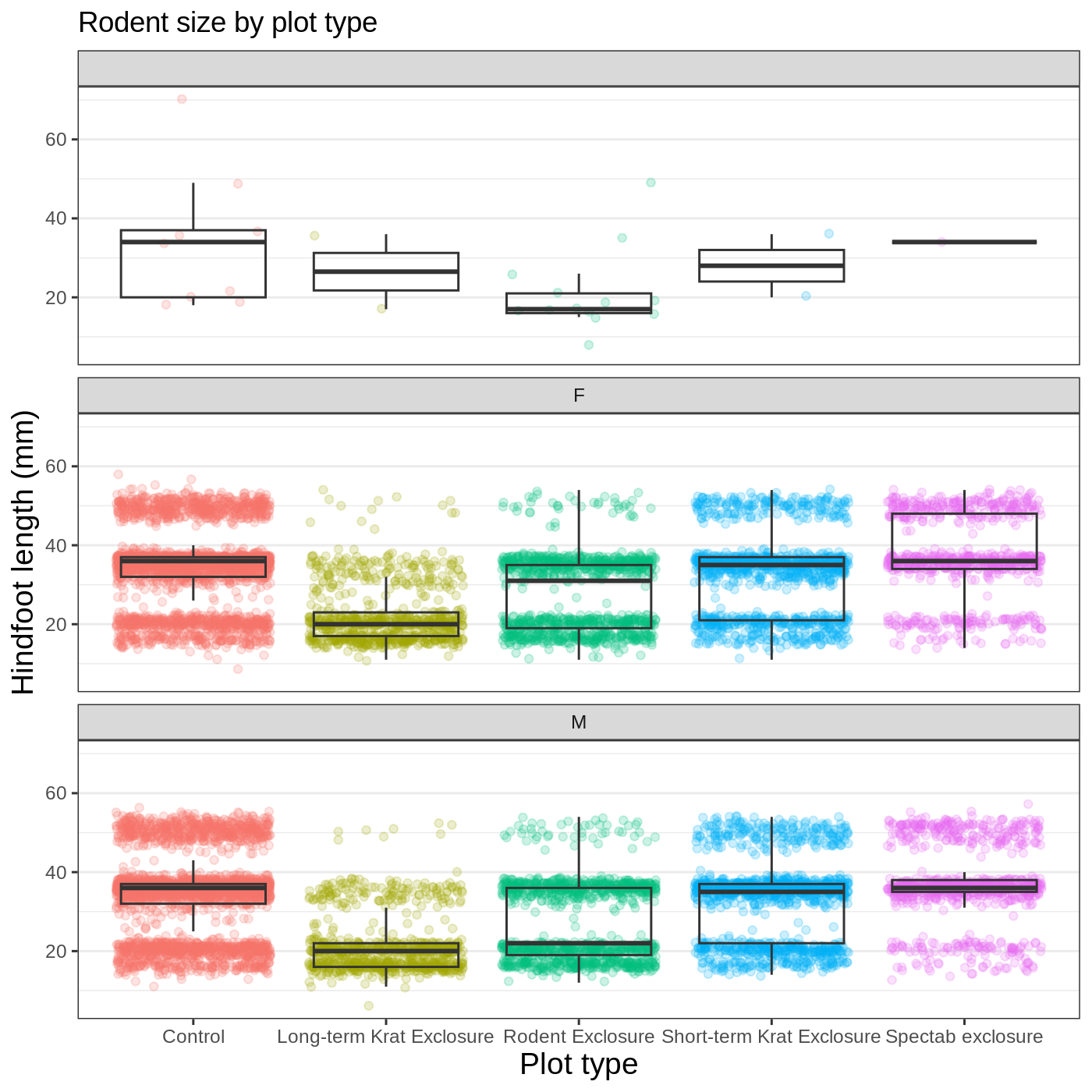
Callout
Faceting comes in handy in many scenarios. It can be useful when:
- a categorical variable has too many levels to differentiate by color (such as a dataset with 20 countries)
- your data overlap heavily, obscuring categories
- you want to show more than 3 variables at once
- you want to see each category in isolation while allowing for general comparisons between categories
Exporting plots
Once we are happy with our final plot, we can assign the whole thing
to a new object, which we can call finalplot.
R
finalplot <- myplot +
theme_bw() +
theme(axis.title = element_text(size = 14),
legend.position = "none",
panel.grid.major.x = element_blank()) +
labs(title = "Rodent size by plot type",
x = "Plot type",
y = "Hindfoot length (mm)",
color = "Plot type") +
facet_wrap(vars(sex), ncol = 1)
After this, we can run ggsave() to save our plot. The
first argument we give is the path to the file we want to save,
including the correct file extension. This code will make an image
called rodent_size_plots.jpg in the images/
folder of our current project. We are making a .jpg, but
you can save .pdf, .tiff, and other file
formats. Next, we tell it the name of the plot object we want to save.
We can also specify things like the width and height of the plot in
inches.
R
ggsave(filename = "images/rodent_size_plots.jpg", plot = finalplot,
height = 6, width = 8)
Challenge 4: Make your own plot
Try making your own plot! You can run str(complete_old)
or ?complete_old to explore variables you might use in your
new plot. Feel free to use variables we have already seen, or some we
haven’t explored yet.
Here are a couple ideas to get you started:
- make a histogram of one of the numeric variables
- try using a different color
scale_ - try changing the size of points or thickness of lines in a
geom
Key Points
- the
ggplot()function initiates a plot, andgeom_functions add representations of your data - use
aes()when mapping a variable from the data to a part of the plot - use
scale_functions to modify the scales used to represent variables - use premade
theme_functions to broadly change appearance, and thetheme()function to fine-tune - start simple and build your plots iteratively
Content from Exploring and understanding data
Last updated on 2024-07-23 | Edit this page
Overview
Questions
- How does R store and represent data?
Objectives
- Explore the structure and content of data.frames
- Understand vector types and missing data
- Use vectors as function arguments
- Create and convert factors
- Understand how R assigns values to objects
Setup
R
library(tidyverse)
library(ratdat)
The data.frame
We just spent quite a bit of time learning how to create
visualizations from the complete_old data, but we did not
talk much about what this complete_old
thing is. It’s important to understand how R thinks about, represents,
and stores data in order for us to have a productive working
relationship with R.
The complete_old data is stored in R as a
data.frame, which is the most common way that R
represents tabular data (data that can be stored in a table format, like
a spreadsheet). We can check what complete_old is by using
the class() function:
R
class(complete_old)
OUTPUT
[1] "tbl_df" "tbl" "data.frame"We can view the first few rows with the head() function,
and the last few rows with the tail() function:
R
head(complete_old)
OUTPUT
# A tibble: 6 × 13
record_id month day year plot_id species_id sex hindfoot_length weight
<int> <int> <int> <int> <int> <chr> <chr> <int> <int>
1 1 7 16 1977 2 NL M 32 NA
2 2 7 16 1977 3 NL M 33 NA
3 3 7 16 1977 2 DM F 37 NA
4 4 7 16 1977 7 DM M 36 NA
5 5 7 16 1977 3 DM M 35 NA
6 6 7 16 1977 1 PF M 14 NA
# ℹ 4 more variables: genus <chr>, species <chr>, taxa <chr>, plot_type <chr>R
tail(complete_old)
OUTPUT
# A tibble: 6 × 13
record_id month day year plot_id species_id sex hindfoot_length weight
<int> <int> <int> <int> <int> <chr> <chr> <int> <int>
1 16873 12 5 1989 8 DO M 37 51
2 16874 12 5 1989 16 RM F 18 15
3 16875 12 5 1989 5 RM M 17 9
4 16876 12 5 1989 4 DM M 37 31
5 16877 12 5 1989 11 DM M 37 50
6 16878 12 5 1989 8 DM F 37 42
# ℹ 4 more variables: genus <chr>, species <chr>, taxa <chr>, plot_type <chr>We used these functions with just one argument, the object
complete_old, and we didn’t give the argument a name, like
we often did with ggplot2. In R, a function’s arguments
come in a particular order, and if you put them in the correct order,
you don’t need to name them. In this case, the name of the argument is
x, so we can name it if we want, but since we know it’s the
first argument, we don’t need to.
To learn more about a function, you can type a ? in
front of the name of the function, which will bring up the official
documentation for that function:
R
?head
Callout
Function documentation is written by the authors of the functions, so they can vary pretty widely in their style and readability. The first section, Description, gives you a concise description of what the function does, but it may not always be enough. The Arguments section defines all the arguments for the function and is usually worth reading thoroughly. Finally, the Examples section at the end will often have some helpful examples that you can run to get a sense of what the function is doing.
Another great source of information is package
vignettes. Many packages have vignettes, which are like
tutorials that introduce the package, specific functions, or general
methods. You can run vignette(package = "package_name") to
see a list of vignettes in that package. Once you have a name, you can
run vignette("vignette_name", "package_name") to view that
vignette. You can also use a web browser to go to
https://cran.r-project.org/web/packages/package_name/vignettes/
where you will find a list of links to each vignette. Some packages will
have their own websites, which often have nicely formatted vignettes and
tutorials.
Finally, learning to search for help is probably the most useful
skill for any R user. The key skill is figuring out what you should
actually search for. It’s often a good idea to start your search with
R or R programming. If you have the name of a
package you want to use, start with R package_name.
Many of the answers you find will be from a website called Stack
Overflow, where people ask programming questions and others provide
answers. It is generally poor form to ask duplicate questions, so before
you decide to post your own, do some thorough searching to see if it has
been answered before (it likely has). If you do decide to post a
question on Stack Overflow, or any other help forum, you will want to
create a reproducible example or
reprex. If you are asking a complicated question
requiring your own data and a whole bunch of code, people probably won’t
be able or willing to help you. However, if you can hone in on the
specific thing you want help with, and create a minimal example using
smaller, fake data, it will be much easier for others to help you. If
you search how to make a reproducible example in R, you
will find some great resources to help you out.
Some arguments are optional. For example, the n argument
in head() specifies the number of rows to print. It
defaults to 6, but we can override that by specifying a different
number:
R
head(complete_old, n = 10)
OUTPUT
# A tibble: 10 × 13
record_id month day year plot_id species_id sex hindfoot_length weight
<int> <int> <int> <int> <int> <chr> <chr> <int> <int>
1 1 7 16 1977 2 NL M 32 NA
2 2 7 16 1977 3 NL M 33 NA
3 3 7 16 1977 2 DM F 37 NA
4 4 7 16 1977 7 DM M 36 NA
5 5 7 16 1977 3 DM M 35 NA
6 6 7 16 1977 1 PF M 14 NA
7 7 7 16 1977 2 PE F NA NA
8 8 7 16 1977 1 DM M 37 NA
9 9 7 16 1977 1 DM F 34 NA
10 10 7 16 1977 6 PF F 20 NA
# ℹ 4 more variables: genus <chr>, species <chr>, taxa <chr>, plot_type <chr>If we order them correctly, we don’t have to name either:
R
head(complete_old, 10)
OUTPUT
# A tibble: 10 × 13
record_id month day year plot_id species_id sex hindfoot_length weight
<int> <int> <int> <int> <int> <chr> <chr> <int> <int>
1 1 7 16 1977 2 NL M 32 NA
2 2 7 16 1977 3 NL M 33 NA
3 3 7 16 1977 2 DM F 37 NA
4 4 7 16 1977 7 DM M 36 NA
5 5 7 16 1977 3 DM M 35 NA
6 6 7 16 1977 1 PF M 14 NA
7 7 7 16 1977 2 PE F NA NA
8 8 7 16 1977 1 DM M 37 NA
9 9 7 16 1977 1 DM F 34 NA
10 10 7 16 1977 6 PF F 20 NA
# ℹ 4 more variables: genus <chr>, species <chr>, taxa <chr>, plot_type <chr>Additionally, if we name them, we can put them in any order we want:
R
head(n = 10, x = complete_old)
OUTPUT
# A tibble: 10 × 13
record_id month day year plot_id species_id sex hindfoot_length weight
<int> <int> <int> <int> <int> <chr> <chr> <int> <int>
1 1 7 16 1977 2 NL M 32 NA
2 2 7 16 1977 3 NL M 33 NA
3 3 7 16 1977 2 DM F 37 NA
4 4 7 16 1977 7 DM M 36 NA
5 5 7 16 1977 3 DM M 35 NA
6 6 7 16 1977 1 PF M 14 NA
7 7 7 16 1977 2 PE F NA NA
8 8 7 16 1977 1 DM M 37 NA
9 9 7 16 1977 1 DM F 34 NA
10 10 7 16 1977 6 PF F 20 NA
# ℹ 4 more variables: genus <chr>, species <chr>, taxa <chr>, plot_type <chr>Generally, it’s good practice to start with the required arguments, like the data.frame whose rows you want to see, and then to name the optional arguments. If you are ever unsure, it never hurts to explicitly name an argument.
Let’s get back to investigating our complete_old
data.frame. We can get some useful summaries of each variable using the
summary() function:
R
summary(complete_old)
OUTPUT
record_id month day year plot_id
Min. : 1 Min. : 1.000 Min. : 1.0 Min. :1977 Min. : 1.00
1st Qu.: 4220 1st Qu.: 3.000 1st Qu.: 9.0 1st Qu.:1981 1st Qu.: 5.00
Median : 8440 Median : 6.000 Median :15.0 Median :1983 Median :11.00
Mean : 8440 Mean : 6.382 Mean :15.6 Mean :1984 Mean :11.47
3rd Qu.:12659 3rd Qu.: 9.000 3rd Qu.:23.0 3rd Qu.:1987 3rd Qu.:17.00
Max. :16878 Max. :12.000 Max. :31.0 Max. :1989 Max. :24.00
species_id sex hindfoot_length weight
Length:16878 Length:16878 Min. : 6.00 Min. : 4.00
Class :character Class :character 1st Qu.:21.00 1st Qu.: 24.00
Mode :character Mode :character Median :35.00 Median : 42.00
Mean :31.98 Mean : 53.22
3rd Qu.:37.00 3rd Qu.: 53.00
Max. :70.00 Max. :278.00
NA's :2733 NA's :1692
genus species taxa plot_type
Length:16878 Length:16878 Length:16878 Length:16878
Class :character Class :character Class :character Class :character
Mode :character Mode :character Mode :character Mode :character
And, as we have already done, we can use str() to look
at the structure of an object:
R
str(complete_old)
OUTPUT
tibble [16,878 × 13] (S3: tbl_df/tbl/data.frame)
$ record_id : int [1:16878] 1 2 3 4 5 6 7 8 9 10 ...
$ month : int [1:16878] 7 7 7 7 7 7 7 7 7 7 ...
$ day : int [1:16878] 16 16 16 16 16 16 16 16 16 16 ...
$ year : int [1:16878] 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977 ...
$ plot_id : int [1:16878] 2 3 2 7 3 1 2 1 1 6 ...
$ species_id : chr [1:16878] "NL" "NL" "DM" "DM" ...
$ sex : chr [1:16878] "M" "M" "F" "M" ...
$ hindfoot_length: int [1:16878] 32 33 37 36 35 14 NA 37 34 20 ...
$ weight : int [1:16878] NA NA NA NA NA NA NA NA NA NA ...
$ genus : chr [1:16878] "Neotoma" "Neotoma" "Dipodomys" "Dipodomys" ...
$ species : chr [1:16878] "albigula" "albigula" "merriami" "merriami" ...
$ taxa : chr [1:16878] "Rodent" "Rodent" "Rodent" "Rodent" ...
$ plot_type : chr [1:16878] "Control" "Long-term Krat Exclosure" "Control" "Rodent Exclosure" ...We get quite a bit of useful information here. First, we are told that we have a data.frame of 16878 observations, or rows, and 13 variables, or columns.
Next, we get a bit of information on each variable, including its
type (int or chr) and a quick peek at the
first 10 values. You might ask why there is a $ in front of
each variable. This is because the $ is an operator that
allows us to select individual columns from a data.frame.
The $ operator also allows you to use tab-completion to
quickly select which variable you want from a given data.frame. For
example, to get the year variable, we can type
complete_old$ and then hit Tab. We get a list of
the variables that we can move through with up and down arrow keys. Hit
Enter when you reach year, which should finish
this code:
R
complete_old$year
OUTPUT
[1] 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977
[16] 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977
[31] 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977
[46] 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977
[61] 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977
[76] 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977
[91] 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977
[ reached getOption("max.print") -- omitted 16778 entries ]What we get back is a whole bunch of numbers, the entries in the
year column printed out in order.
Vectors: the building block of data
You might have noticed that our last result looked different from
when we printed out the complete_old data.frame itself.
That’s because it is not a data.frame, it is a vector.
A vector is a 1-dimensional series of values, in this case a vector of
numbers representing years.
Data.frames are made up of vectors; each column in a data.frame is a vector. Vectors are the basic building blocks of all data in R. Basically, everything in R is a vector, a bunch of vectors stitched together in some way, or a function. Understanding how vectors work is crucial to understanding how R treats data, so we will spend some time learning about them.
There are 4 main types of vectors (also known as atomic vectors):
"character"for strings of characters, like ourgenusorsexcolumns. Each entry in a character vector is wrapped in quotes. In other programming languages, this type of data may be referred to as “strings”."integer"for integers. All the numeric values incomplete_oldare integers. You may sometimes see integers represented like2Lor20L. TheLindicates to R that it is an integer, instead of the next data type,"numeric"."numeric", aka"double", vectors can contain numbers including decimals. Other languages may refer to these as “float” or “floating point” numbers."logical"forTRUEandFALSE, which can also be represented asTandF. In other contexts, these may be referred to as “Boolean” data.
Vectors can only be of a single type. Since each
column in a data.frame is a vector, this means an accidental character
following a number, like 29, can change the type of the
whole vector. Mixing up vector types is one of the most common mistakes
in R, and it can be tricky to figure out. It’s often very useful to
check the types of vectors.
To create a vector from scratch, we can use the c()
function, putting values inside, separated by commas.
R
c(1, 2, 5, 12, 4)
OUTPUT
[1] 1 2 5 12 4As you can see, those values get printed out in the console, just
like with complete_old$year. To store this vector so we can
continue to work with it, we need to assign it to an object.
R
num <- c(1, 2, 5, 12, 4)
You can check what kind of object num is with the
class() function.
R
class(num)
OUTPUT
[1] "numeric"We see that num is a numeric vector.
Let’s try making a character vector:
R
char <- c("apple", "pear", "grape")
class(char)
OUTPUT
[1] "character"Remember that each entry, like "apple", needs to be
surrounded by quotes, and entries are separated with commas. If you do
something like "apple, pear, grape", you will have only a
single entry containing that whole string.
Finally, let’s make a logical vector:
R
logi <- c(TRUE, FALSE, TRUE, TRUE)
class(logi)
OUTPUT
[1] "logical"Challenge 1: Coercion
Since vectors can only hold one type of data, something has to be done when we try to combine different types of data into one vector.
- What type will each of these vectors be? Try to guess without
running any code at first, then run the code and use
class()to verify your answers.
R
num_logi <- c(1, 4, 6, TRUE)
num_char <- c(1, 3, "10", 6)
char_logi <- c("a", "b", TRUE)
tricky <- c("a", "b", "1", FALSE)
R
class(num_logi)
OUTPUT
[1] "numeric"R
class(num_char)
OUTPUT
[1] "character"R
class(char_logi)
OUTPUT
[1] "character"R
class(tricky)
OUTPUT
[1] "character"R will automatically convert values in a vector so that they are all the same type, a process called coercion.
R
combined_logical
OUTPUT
[1] "1" "4" "6" "1" "a" "b" "TRUE"R
class(combined_logical)
OUTPUT
[1] "character"Only one value is "TRUE". Coercion happens when each
vector is created, so the TRUE in num_logi
becomes a 1, while the TRUE in
char_logi becomes "TRUE". When these two
vectors are combined, R doesn’t remember that the 1 in
num_logi used to be a TRUE, it will just
coerce the 1 to "1".
Challenge 1: Coercion (continued)
- Now that you’ve seen a few examples of coercion, you might have started to see that there are some rules about how types get converted. There is a hierarchy to coercion. Can you draw a diagram that represents the hierarchy of what types get converted to other types?
logical → integer → numeric → character
Logical vectors can only take on two values: TRUE or
FALSE. Integer vectors can only contain integers, so
TRUE and FALSE can be coerced to
1 and 0. Numeric vectors can contain numbers
with decimals, so integers can be coerced from, say, 6 to
6.0 (though R will still display a numeric 6
as 6.). Finally, any string of characters can be
represented as a character vector, so any of the other types can be
coerced to a character vector.
Coercion is not something you will often do intentionally; rather,
when combining vectors or reading data into R, a stray character that
you missed may change an entire numeric vector into a character vector.
It is a good idea to check the class() of your results
frequently, particularly if you are running into confusing error
messages.
Missing data
One of the great things about R is how it handles missing data, which
can be tricky in other programming languages. R represents missing data
as NA, without quotes, in vectors of any type. Let’s make a
numeric vector with an NA value:
R
weights <- c(25, 34, 12, NA, 42)
R doesn’t make assumptions about how you want to handle missing data,
so if we pass this vector to a numeric function like min(),
it won’t know what to do, so it returns NA:
R
min(weights)
OUTPUT
[1] NAThis is a very good thing, since we won’t accidentally forget to consider our missing data. If we decide to exclude our missing values, many basic math functions have an argument to remove them:
R
min(weights, na.rm = TRUE)
OUTPUT
[1] 12Vectors as arguments
A common reason to create a vector from scratch is to use in a
function argument. The quantile() function will calculate a
quantile for a given vector of numeric values. We set the quantile using
the probs argument. We also need to set
na.rm = TRUE, since there are NA values in the
weight column.
R
quantile(complete_old$weight, probs = 0.25, na.rm = TRUE)
OUTPUT
25%
24 Now we get back the 25% quantile value for weights. However, we often
want to know more than one quantile. Luckily, the probs
argument is vectorized, meaning it can take a whole
vector of values. Let’s try getting the 25%, 50% (median), and 75%
quantiles all at once.
R
quantile(complete_old$weight, probs = c(0.25, 0.5, 0.75), na.rm = TRUE)
OUTPUT
25% 50% 75%
24 42 53 While the c() function is very flexible, it doesn’t
necessarily scale well. If you want to generate a long vector from
scratch, you probably don’t want to type everything out manually. There
are a few functions that can help generate vectors.
First, putting : between two numbers will generate a
vector of integers starting with the first number and ending with the
last. The seq() function allows you to generate similar
sequences, but changing by any amount.
R
# generates a sequence of integers
1:10
OUTPUT
[1] 1 2 3 4 5 6 7 8 9 10R
# with seq() you can generate sequences with a combination of:
# from: starting value
# to: ending value
# by: how much should each entry increase
# length.out: how long should the resulting vector be
seq(from = 0, to = 1, by = 0.1)
OUTPUT
[1] 0.0 0.1 0.2 0.3 0.4 0.5 0.6 0.7 0.8 0.9 1.0R
seq(from = 0, to = 1, length.out = 50)
OUTPUT
[1] 0.00000000 0.02040816 0.04081633 0.06122449 0.08163265 0.10204082
[7] 0.12244898 0.14285714 0.16326531 0.18367347 0.20408163 0.22448980
[13] 0.24489796 0.26530612 0.28571429 0.30612245 0.32653061 0.34693878
[19] 0.36734694 0.38775510 0.40816327 0.42857143 0.44897959 0.46938776
[25] 0.48979592 0.51020408 0.53061224 0.55102041 0.57142857 0.59183673
[31] 0.61224490 0.63265306 0.65306122 0.67346939 0.69387755 0.71428571
[37] 0.73469388 0.75510204 0.77551020 0.79591837 0.81632653 0.83673469
[43] 0.85714286 0.87755102 0.89795918 0.91836735 0.93877551 0.95918367
[49] 0.97959184 1.00000000R
seq(from = 0, by = 0.01, length.out = 20)
OUTPUT
[1] 0.00 0.01 0.02 0.03 0.04 0.05 0.06 0.07 0.08 0.09 0.10 0.11 0.12 0.13 0.14
[16] 0.15 0.16 0.17 0.18 0.19Finally, the rep() function allows you to repeat a
value, or even a whole vector, as many times as you want, and works with
any type of vector.
R
# repeats "a" 12 times
rep("a", times = 12)
OUTPUT
[1] "a" "a" "a" "a" "a" "a" "a" "a" "a" "a" "a" "a"R
# repeats this whole sequence 4 times
rep(c("a", "b", "c"), times = 4)
OUTPUT
[1] "a" "b" "c" "a" "b" "c" "a" "b" "c" "a" "b" "c"R
# repeats each value 4 times
rep(1:10, each = 4)
OUTPUT
[1] 1 1 1 1 2 2 2 2 3 3 3 3 4 4 4 4 5 5 5 5 6 6 6 6 7
[26] 7 7 7 8 8 8 8 9 9 9 9 10 10 10 10R
rep(-3:3, 3)
OUTPUT
[1] -3 -2 -1 0 1 2 3 -3 -2 -1 0 1 2 3 -3 -2 -1 0 1 2 3R
# this also works
rep(seq(from = -3, to = 3, by = 1), 3)
OUTPUT
[1] -3 -2 -1 0 1 2 3 -3 -2 -1 0 1 2 3 -3 -2 -1 0 1 2 3R
# you might also store the sequence as an intermediate vector
my_seq <- seq(from = -3, to = 3, by = 1)
rep(my_seq, 3)
OUTPUT
[1] -3 -2 -1 0 1 2 3 -3 -2 -1 0 1 2 3 -3 -2 -1 0 1 2 3R
quantile(complete_old$hindfoot_length,
probs = seq(from = 0, to = 1, by = 0.05),
na.rm = T)
OUTPUT
0% 5% 10% 15% 20% 25% 30% 35% 40% 45% 50% 55% 60% 65% 70% 75%
6 16 17 19 20 21 22 31 33 34 35 35 36 36 36 37
80% 85% 90% 95% 100%
37 39 49 51 70 Building with vectors
We have now seen vectors in a few different forms: as columns in a data.frame and as single vectors. However, they can be manipulated into lots of other shapes and forms. Some other common forms are:
- matrices
- 2-dimensional numeric representations
- arrays
- many-dimensional numeric
- lists
- lists are very flexible ways to store vectors
- a list can contain vectors of many different types and lengths
- an entry in a list can be another list, so lists can get deeply nested
- a data.frame is a type of list where each column is an individual vector and each vector has to be the same length, since a data.frame has an entry in every column for each row
- factors
- a way to represent categorical data
- factors can be ordered or unordered
- they often look like character vectors, but behave differently
- under the hood, they are integers with character labels, called levels, for each integer
Factors
We will spend a bit more time talking about factors, since they are
often a challenging type of data to work with. We can create a factor
from scratch by putting a character vector made using c()
into the factor() function:
R
sex <- factor(c("male", "female", "female", "male", "female", NA))
sex
OUTPUT
[1] male female female male female <NA>
Levels: female maleWe can inspect the levels of the factor using the
levels() function:
R
levels(sex)
OUTPUT
[1] "female" "male" The forcats package from the
tidyverse has a lot of convenient functions for working
with factors. We will show you a few common operations, but the
forcats package has many more useful functions.
R
library(forcats)
# change the order of the levels
fct_relevel(sex, c("male", "female"))
OUTPUT
[1] male female female male female <NA>
Levels: male femaleR
# change the names of the levels
fct_recode(sex, "M" = "male", "F" = "female")
OUTPUT
[1] M F F M F <NA>
Levels: F MR
# turn NAs into an actual factor level (useful for including NAs in plots)
fct_na_value_to_level(sex, "(Missing)")
OUTPUT
[1] male female female male female (Missing)
Levels: female male (Missing)In general, it is a good practice to leave your categorical data as a character vector until you need to use a factor. Here are some reasons you might need a factor:
- Another function requires you to use a factor
- You are plotting categorical data and want to control the ordering of categories in the plot
Since factors can behave differently from character vectors, it is always a good idea to check what type of data you’re working with. You might use a new function for the first time and be confused by the results, only to realize later that it produced a factor as an output, when you thought it was a character vector.
It is fairly straightforward to convert a factor to a character vector:
R
as.character(sex)
OUTPUT
[1] "male" "female" "female" "male" "female" NA However, you need to be careful if you’re somehow working with a factor that has numbers as its levels:
R
f_num <- factor(c(1990, 1983, 1977, 1998, 1990))
# this will pull out the underlying integers, not the levels
as.numeric(f_num)
OUTPUT
[1] 3 2 1 4 3R
# if we first convert to characters, we can then convert to numbers
as.numeric(as.character(f_num))
OUTPUT
[1] 1990 1983 1977 1998 1990Assignment, objects, and values
We’ve already created quite a few objects in R using the
<- assignment arrow, but there are a few finer details
worth talking about. First, let’s start with a quick challenge.
R
x <- 5
y <- x
x <- 10
y
OUTPUT
[1] 5Understanding what’s going on here will help you avoid a lot of
confusion when working in R. When we assign something to an object, the
first thing that happens is the righthand side gets evaluated.
The same thing happens when you run something in the console: if you
type x into the console and hit Enter, R returns
the value of x. So when we first ran the line
y <- x, x first gets evaluated to the value
of 5, and this gets assigned to y. The objects
x and y are not actually linked to each other
in any way, so when we change the value of x to
10, y is unaffected.
This also means you can run multiple nested operations, store intermediate values as separate objects, or overwrite values:
R
x <- 5
# first, x gets evaluated to 5
# then 5/2 gets evaluated to 2.5
# then sqrt(2.5) is evaluated
sqrt(x/2)
OUTPUT
[1] 1.581139R
# we can also store the evaluated value of x/2
# in an object y before passing it to sqrt()
y <- x/2
sqrt(y)
OUTPUT
[1] 1.581139R
# first, the x on the righthand side gets evaluated to 5
# then 5 gets squared
# then the resulting value is assigned to the object x
x <- x^2
x
OUTPUT
[1] 25You will be naming a of objects in R, and there are a few common naming rules and conventions:
- make names clear without being too long
-
wkgis probably too short -
weight_in_kilogramsis probably too long -
weight_kgis good
-
- names cannot start with a number
- names are case sensitive
- you cannot use the names of fundamental functions in R, like
if,else, orfor- in general, avoid using names of common functions like
c,mean, etc.
- in general, avoid using names of common functions like
- avoid dots
.in names, as they have a special meaning in R, and may be confusing to others - two common formats are
snake_caseandcamelCase - be consistent, at least within a script, ideally within a whole project
- you can use a style guide like Google’s or tidyverse’s
Key Points
- functions like
head(),str(), andsummary()are useful for exploring data.frames - most things in R are vectors, vectors stitched together, or functions
- make sure to use
class()to check vector types, especially when using new functions - factors can be useful, but behave differently from character vectors
Content from Working with data
Last updated on 2024-07-23 | Edit this page
Overview
Questions
- How do you manipulate tabular data in R?
Objectives
- Import CSV data into R.
- Understand the difference between base R and
tidyverseapproaches. - Subset rows and columns of data.frames.
- Use pipes to link steps together into pipelines.
- Create new data.frame columns using existing columns.
- Utilize the concept of split-apply-combine data analysis.
- Reshape data between wide and long formats.
- Export data to a CSV file.
R
library(tidyverse)
Importing data
Up until this point, we have been working with the
complete_old dataframe contained in the ratdat
package. However, you typically won’t access data from an R package; it
is much more common to access data files stored somewhere on your
computer. We are going to download a CSV file containing the surveys
data to our computer, which we will then read into R.
Click this link to download the file: https://datacarpentry.org/R-ecology-lesson/data/cleaned/surveys_complete_77_89.csv.
You will be prompted to save the file on your computer somewhere.
Save it inside the cleaned data folder, which is in the
data folder in your R-Ecology-Workshop folder.
Once it’s inside our project, we will be able to point R towards it.
File paths
When we reference other files from an R script, we need to give R
precise instructions on where those files are. We do that using
something called a file path. It looks something like
this: "Documents/Manuscripts/Chapter_2.txt". This path
would tell your computer how to get from whatever folder contains the
Documents folder all the way to the .txt
file.
There are two kinds of paths: absolute and
relative. Absolute paths are specific to a particular
computer, whereas relative paths are relative to a certain folder.
Because we are keeping all of our work in the
R-Ecology-Workshop folder, all of our paths can be relative
to this folder.
Now, let’s read our CSV file into R and store it in an object named
surveys. We will use the read_csv function
from the tidyverse’s readr package, and the
argument we give will be the relative path to the CSV
file.
R
surveys <- read_csv("data/cleaned/surveys_complete_77_89.csv")
OUTPUT
Rows: 16878 Columns: 13
── Column specification ────────────────────────────────────────────────────────
Delimiter: ","
chr (6): species_id, sex, genus, species, taxa, plot_type
dbl (7): record_id, month, day, year, plot_id, hindfoot_length, weight
ℹ Use `spec()` to retrieve the full column specification for this data.
ℹ Specify the column types or set `show_col_types = FALSE` to quiet this message.Callout
Typing out paths can be error prone, so we can utilize a keyboard
shortcut. Inside the parentheses of read_csv(), type out a
pair of quotes and put your cursor between them. Then hit
Tab. A small menu showing your folders and files should show
up. You can use the ↑ and ↓ keys to move through
the options, or start typing to narrow them down. You can hit
Enter to select a file or folder, and hit Tab
again to continue building the file path. This might take a bit of
getting used to, but once you get the hang of it, it will speed up
writing file paths and reduce the number of mistakes you make.
You may have noticed a bit of feedback from R when you ran the last line of code. We got some useful information about the CSV file we read in. We can see:
- the number of rows and columns
- the delimiter of the file, which is how values are
separated, a comma
"," - a set of columns that were parsed as various vector
types
- the file has 6 character columns and 7 numeric columns
- we can see the names of the columns for each type
When working with the output of a new function, it’s often a good
idea to check the class():
R
class(surveys)
OUTPUT
[1] "spec_tbl_df" "tbl_df" "tbl" "data.frame" Whoa! What is this thing? It has multiple classes? Well, it’s called
a tibble, and it is the tidyverse version of a
data.frame. It is a data.frame, but with some added perks. It
prints out a little more nicely, it highlights NA values
and negative values in red, and it will generally communicate with you
more (in terms of warnings and errors, which is a good thing).
Callout
tidyverse vs. base R
As we begin to delve more deeply into the tidyverse, we
should briefly pause to mention some of the reasons for focusing on the
tidyverse set of tools. In R, there are often many ways to
get a job done, and there are other approaches that can accomplish tasks
similar to the tidyverse.
The phrase base R is used to refer to approaches
that utilize functions contained in R’s default packages. We have
already used some base R functions, such as str(),
head(), and mean(), and we will be using more
scattered throughout this lesson. However, there are some key base R
approaches we will not be teaching. These include square bracket
subsetting and base plotting. You may come across code written by other
people that looks like surveys[1:10, 2] or
plot(surveys$weight, surveys$hindfoot_length), which are
base R commands. If you’re interested in learning more about these
approaches, you can check out other Carpentries lessons like the Software
Carpentry Programming with R lesson.
We choose to teach the tidyverse set of packages because
they share a similar syntax and philosophy, making them consistent and
producing highly readable code. They are also very flexible and
powerful, with a growing number of packages designed according to
similar principles and to work well with the rest of the packages. The
tidyverse packages tend to have very clear documentation
and wide array of learning materials that tend to be written with novice
users in mind. Finally, the tidyverse has only continued to
grow, and has strong support from RStudio, which implies that these
approaches will be relevant into the future.
Manipulating data
One of the most important skills for working with data in R is the
ability to manipulate, modify, and reshape data. The dplyr
and tidyr packages in the tidyverse provide a
series of powerful functions for many common data manipulation
tasks.
We’ll start off with two of the most commonly used dplyr
functions: select(), which selects certain columns of a
data.frame, and filter(), which filters out rows according
to certain criteria.
select()
To use the select() function, the first argument is the
name of the data.frame, and the rest of the arguments are
unquoted names of the columns you want:
R
select(surveys, plot_id, species_id, hindfoot_length)
OUTPUT
# A tibble: 16,878 × 3
plot_id species_id hindfoot_length
<dbl> <chr> <dbl>
1 2 NL 32
2 3 NL 33
3 2 DM 37
4 7 DM 36
5 3 DM 35
6 1 PF 14
7 2 PE NA
8 1 DM 37
9 1 DM 34
10 6 PF 20
# ℹ 16,868 more rowsThe columns are arranged in the order we specified inside
select().
To select all columns except specific columns, put a -
in front of the column you want to exclude:
R
select(surveys, -record_id, -year)
OUTPUT
# A tibble: 16,878 × 11
month day plot_id species_id sex hindfoot_length weight genus species
<dbl> <dbl> <dbl> <chr> <chr> <dbl> <dbl> <chr> <chr>
1 7 16 2 NL M 32 NA Neotoma albigu…
2 7 16 3 NL M 33 NA Neotoma albigu…
3 7 16 2 DM F 37 NA Dipodomys merria…
4 7 16 7 DM M 36 NA Dipodomys merria…
5 7 16 3 DM M 35 NA Dipodomys merria…
6 7 16 1 PF M 14 NA Perognat… flavus
7 7 16 2 PE F NA NA Peromysc… eremic…
8 7 16 1 DM M 37 NA Dipodomys merria…
9 7 16 1 DM F 34 NA Dipodomys merria…
10 7 16 6 PF F 20 NA Perognat… flavus
# ℹ 16,868 more rows
# ℹ 2 more variables: taxa <chr>, plot_type <chr>select() also works with numeric vectors for the order
of the columns. To select the 3rd, 4th, 5th, and 10th columns, we could
run the following code:
R
select(surveys, c(3:5, 10))
OUTPUT
# A tibble: 16,878 × 4
day year plot_id genus
<dbl> <dbl> <dbl> <chr>
1 16 1977 2 Neotoma
2 16 1977 3 Neotoma
3 16 1977 2 Dipodomys
4 16 1977 7 Dipodomys
5 16 1977 3 Dipodomys
6 16 1977 1 Perognathus
7 16 1977 2 Peromyscus
8 16 1977 1 Dipodomys
9 16 1977 1 Dipodomys
10 16 1977 6 Perognathus
# ℹ 16,868 more rowsYou should be careful when using this method, since you are being less explicit about which columns you want. However, it can be useful if you have a data.frame with many columns and you don’t want to type out too many names.
Finally, you can select columns based on whether they match a certain
criteria by using the where() function. If we want all
numeric columns, we can ask to select all the columns
where the class is numeric:
R
select(surveys, where(is.numeric))
OUTPUT
# A tibble: 16,878 × 7
record_id month day year plot_id hindfoot_length weight
<dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
1 1 7 16 1977 2 32 NA
2 2 7 16 1977 3 33 NA
3 3 7 16 1977 2 37 NA
4 4 7 16 1977 7 36 NA
5 5 7 16 1977 3 35 NA
6 6 7 16 1977 1 14 NA
7 7 7 16 1977 2 NA NA
8 8 7 16 1977 1 37 NA
9 9 7 16 1977 1 34 NA
10 10 7 16 1977 6 20 NA
# ℹ 16,868 more rowsInstead of giving names or positions of columns, we instead pass the
where() function with the name of another function inside
it, in this case is.numeric(), and we get all the columns
for which that function returns TRUE.
We can use this to select any columns that have any NA
values in them:
R
select(surveys, where(anyNA))
OUTPUT
# A tibble: 16,878 × 7
species_id sex hindfoot_length weight genus species taxa
<chr> <chr> <dbl> <dbl> <chr> <chr> <chr>
1 NL M 32 NA Neotoma albigula Rodent
2 NL M 33 NA Neotoma albigula Rodent
3 DM F 37 NA Dipodomys merriami Rodent
4 DM M 36 NA Dipodomys merriami Rodent
5 DM M 35 NA Dipodomys merriami Rodent
6 PF M 14 NA Perognathus flavus Rodent
7 PE F NA NA Peromyscus eremicus Rodent
8 DM M 37 NA Dipodomys merriami Rodent
9 DM F 34 NA Dipodomys merriami Rodent
10 PF F 20 NA Perognathus flavus Rodent
# ℹ 16,868 more rows
filter()
The filter() function is used to select rows that meet
certain criteria. To get all the rows where the value of
year is equal to 1985, we would run the following:
R
filter(surveys, year == 1985)
OUTPUT
# A tibble: 1,438 × 13
record_id month day year plot_id species_id sex hindfoot_length weight
<dbl> <dbl> <dbl> <dbl> <dbl> <chr> <chr> <dbl> <dbl>
1 9790 1 19 1985 16 RM F 16 4
2 9791 1 19 1985 17 OT F 20 16
3 9792 1 19 1985 6 DO M 35 48
4 9793 1 19 1985 12 DO F 35 40
5 9794 1 19 1985 24 RM M 16 4
6 9795 1 19 1985 12 DO M 34 48
7 9796 1 19 1985 6 DM F 37 35
8 9797 1 19 1985 14 DM M 36 45
9 9798 1 19 1985 6 DM F 36 38
10 9799 1 19 1985 19 RM M 16 4
# ℹ 1,428 more rows
# ℹ 4 more variables: genus <chr>, species <chr>, taxa <chr>, plot_type <chr>The == sign means “is equal to”. There are several other
operators we can use: >, >=, <, <=, and != (not equal to).
Another useful operator is %in%, which asks if the value on
the lefthand side is found anywhere in the vector on the righthand side.
For example, to get rows with specific species_id values,
we could run:
R
filter(surveys, species_id %in% c("RM", "DO"))
OUTPUT
# A tibble: 2,835 × 13
record_id month day year plot_id species_id sex hindfoot_length weight
<dbl> <dbl> <dbl> <dbl> <dbl> <chr> <chr> <dbl> <dbl>
1 68 8 19 1977 8 DO F 32 52
2 292 10 17 1977 3 DO F 36 33
3 294 10 17 1977 3 DO F 37 50
4 311 10 17 1977 19 RM M 18 13
5 317 10 17 1977 17 DO F 32 48
6 323 10 17 1977 17 DO F 33 31
7 337 10 18 1977 8 DO F 35 41
8 356 11 12 1977 1 DO F 32 44
9 378 11 12 1977 1 DO M 33 48
10 397 11 13 1977 17 RM F 16 7
# ℹ 2,825 more rows
# ℹ 4 more variables: genus <chr>, species <chr>, taxa <chr>, plot_type <chr>We can also use multiple conditions in one filter()
statement. Here we will get rows with a year less than or equal to 1988
and whose hindfoot length values are not NA. The
! before the is.na() function means “not”.
R
filter(surveys, year <= 1988 & !is.na(hindfoot_length))
OUTPUT
# A tibble: 12,779 × 13
record_id month day year plot_id species_id sex hindfoot_length weight
<dbl> <dbl> <dbl> <dbl> <dbl> <chr> <chr> <dbl> <dbl>
1 1 7 16 1977 2 NL M 32 NA
2 2 7 16 1977 3 NL M 33 NA
3 3 7 16 1977 2 DM F 37 NA
4 4 7 16 1977 7 DM M 36 NA
5 5 7 16 1977 3 DM M 35 NA
6 6 7 16 1977 1 PF M 14 NA
7 8 7 16 1977 1 DM M 37 NA
8 9 7 16 1977 1 DM F 34 NA
9 10 7 16 1977 6 PF F 20 NA
10 11 7 16 1977 5 DS F 53 NA
# ℹ 12,769 more rows
# ℹ 4 more variables: genus <chr>, species <chr>, taxa <chr>, plot_type <chr>R
surveys_filtered <- filter(surveys, year >= 1980 & year <= 1985)
R
surveys_selected <- select(surveys, year, month, species_id, plot_id)
The pipe: %>%
What happens if we want to both select() and
filter() our data? We have a couple options. First, we
could use nested functions:
R
filter(select(surveys, -day), month >= 7)
OUTPUT
# A tibble: 8,244 × 12
record_id month year plot_id species_id sex hindfoot_length weight genus
<dbl> <dbl> <dbl> <dbl> <chr> <chr> <dbl> <dbl> <chr>
1 1 7 1977 2 NL M 32 NA Neotoma
2 2 7 1977 3 NL M 33 NA Neotoma
3 3 7 1977 2 DM F 37 NA Dipodo…
4 4 7 1977 7 DM M 36 NA Dipodo…
5 5 7 1977 3 DM M 35 NA Dipodo…
6 6 7 1977 1 PF M 14 NA Perogn…
7 7 7 1977 2 PE F NA NA Peromy…
8 8 7 1977 1 DM M 37 NA Dipodo…
9 9 7 1977 1 DM F 34 NA Dipodo…
10 10 7 1977 6 PF F 20 NA Perogn…
# ℹ 8,234 more rows
# ℹ 3 more variables: species <chr>, taxa <chr>, plot_type <chr>R will evaluate statements from the inside out. First,
select() will operate on the surveys
data.frame, removing the column day. The resulting
data.frame is then used as the first argument for filter(),
which selects rows with a month greater than or equal to 7.
Nested functions can be very difficult to read with only a few functions, and nearly impossible when many functions are done at once. An alternative approach is to create intermediate objects:
R
surveys_noday <- select(surveys, -day)
filter(surveys_noday, month >= 7)
OUTPUT
# A tibble: 8,244 × 12
record_id month year plot_id species_id sex hindfoot_length weight genus
<dbl> <dbl> <dbl> <dbl> <chr> <chr> <dbl> <dbl> <chr>
1 1 7 1977 2 NL M 32 NA Neotoma
2 2 7 1977 3 NL M 33 NA Neotoma
3 3 7 1977 2 DM F 37 NA Dipodo…
4 4 7 1977 7 DM M 36 NA Dipodo…
5 5 7 1977 3 DM M 35 NA Dipodo…
6 6 7 1977 1 PF M 14 NA Perogn…
7 7 7 1977 2 PE F NA NA Peromy…
8 8 7 1977 1 DM M 37 NA Dipodo…
9 9 7 1977 1 DM F 34 NA Dipodo…
10 10 7 1977 6 PF F 20 NA Perogn…
# ℹ 8,234 more rows
# ℹ 3 more variables: species <chr>, taxa <chr>, plot_type <chr>This approach is easier to read, since we can see the steps in order, but after enough steps, we are left with a cluttered mess of intermediate objects, often with confusing names.
An elegant solution to this problem is an operator called the
pipe, which looks like %>%. You can
insert it by using the keyboard shortcut Shift+Cmd+M (Mac) or
Shift+Ctrl+M (Windows). Here’s how you could use a pipe to
select and filter in one step:
R
surveys %>%
select(-day) %>%
filter(month >= 7)
OUTPUT
# A tibble: 8,244 × 12
record_id month year plot_id species_id sex hindfoot_length weight genus
<dbl> <dbl> <dbl> <dbl> <chr> <chr> <dbl> <dbl> <chr>
1 1 7 1977 2 NL M 32 NA Neotoma
2 2 7 1977 3 NL M 33 NA Neotoma
3 3 7 1977 2 DM F 37 NA Dipodo…
4 4 7 1977 7 DM M 36 NA Dipodo…
5 5 7 1977 3 DM M 35 NA Dipodo…
6 6 7 1977 1 PF M 14 NA Perogn…
7 7 7 1977 2 PE F NA NA Peromy…
8 8 7 1977 1 DM M 37 NA Dipodo…
9 9 7 1977 1 DM F 34 NA Dipodo…
10 10 7 1977 6 PF F 20 NA Perogn…
# ℹ 8,234 more rows
# ℹ 3 more variables: species <chr>, taxa <chr>, plot_type <chr>What it does is take the thing on the lefthand side and insert it as
the first argument of the function on the righthand side. By putting
each of our functions onto a new line, we can build a nice, readable
pipeline. It can be useful to think of this as a little
assembly line for our data. It starts at the top and gets piped into a
select() function, and it comes out modified somewhat. It
then gets sent into the filter() function, where it is
further modified, and then the final product gets printed out to our
console. It can also be helpful to think of %>% as
meaning “and then”. Since many tidyverse functions have
verbs for names, a pipeline can be read like a sentence.
If we want to store this final product as an object, we use an assignment arrow at the start:
R
surveys_sub <- surveys %>%
select(-day) %>%
filter(month >= 7)
A good approach is to build a pipeline step by step prior to assignment. You add functions to the pipeline as you go, with the results printing in the console for you to view. Once you’re satisfied with your final result, go back and add the assignment arrow statement at the start. This approach is very interactive, allowing you to see the results of each step as you build the pipeline, and produces nicely readable code.
R
surveys_1988 <- surveys %>%
filter(year == 1988) %>%
select(record_id, month, species_id)
Make sure to filter() before you select().
You need to use the year column for filtering rows, but it
is discarded in the select() step. You also need to make
sure to use == instead of = when you are
filtering rows where year is equal to 1988.
Making new columns with mutate()
Another common task is creating a new column based on values in existing columns. For example, we could add a new column that has the weight in kilograms instead of grams:
R
surveys %>%
mutate(weight_kg = weight / 1000)
OUTPUT
# A tibble: 16,878 × 14
record_id month day year plot_id species_id sex hindfoot_length weight
<dbl> <dbl> <dbl> <dbl> <dbl> <chr> <chr> <dbl> <dbl>
1 1 7 16 1977 2 NL M 32 NA
2 2 7 16 1977 3 NL M 33 NA
3 3 7 16 1977 2 DM F 37 NA
4 4 7 16 1977 7 DM M 36 NA
5 5 7 16 1977 3 DM M 35 NA
6 6 7 16 1977 1 PF M 14 NA
7 7 7 16 1977 2 PE F NA NA
8 8 7 16 1977 1 DM M 37 NA
9 9 7 16 1977 1 DM F 34 NA
10 10 7 16 1977 6 PF F 20 NA
# ℹ 16,868 more rows
# ℹ 5 more variables: genus <chr>, species <chr>, taxa <chr>, plot_type <chr>,
# weight_kg <dbl>You can create multiple columns in one mutate() call,
and they will get created in the order you write them. This means you
can even reference the first new column in the second new column:
R
surveys %>%
mutate(weight_kg = weight / 1000,
weight_lbs = weight_kg * 2.2)
OUTPUT
# A tibble: 16,878 × 15
record_id month day year plot_id species_id sex hindfoot_length weight
<dbl> <dbl> <dbl> <dbl> <dbl> <chr> <chr> <dbl> <dbl>
1 1 7 16 1977 2 NL M 32 NA
2 2 7 16 1977 3 NL M 33 NA
3 3 7 16 1977 2 DM F 37 NA
4 4 7 16 1977 7 DM M 36 NA
5 5 7 16 1977 3 DM M 35 NA
6 6 7 16 1977 1 PF M 14 NA
7 7 7 16 1977 2 PE F NA NA
8 8 7 16 1977 1 DM M 37 NA
9 9 7 16 1977 1 DM F 34 NA
10 10 7 16 1977 6 PF F 20 NA
# ℹ 16,868 more rows
# ℹ 6 more variables: genus <chr>, species <chr>, taxa <chr>, plot_type <chr>,
# weight_kg <dbl>, weight_lbs <dbl>We can also use multiple columns to create a single column. For example, it’s often good practice to keep the components of a date in separate columns until necessary, as we’ve done here. This is because programs like Excel can do automatic things with dates in a way that is not reproducible and sometimes hard to notice. However, now that we are working in R, we can safely put together a date column.
To put together the columns into something that looks like a date, we
can use the paste() function, which takes arguments of the
items to paste together, as well as the argument sep, which
is the character used to separate the items.
R
surveys %>%
mutate(date = paste(year, month, day, sep = "-"))
OUTPUT
# A tibble: 16,878 × 14
record_id month day year plot_id species_id sex hindfoot_length weight
<dbl> <dbl> <dbl> <dbl> <dbl> <chr> <chr> <dbl> <dbl>
1 1 7 16 1977 2 NL M 32 NA
2 2 7 16 1977 3 NL M 33 NA
3 3 7 16 1977 2 DM F 37 NA
4 4 7 16 1977 7 DM M 36 NA
5 5 7 16 1977 3 DM M 35 NA
6 6 7 16 1977 1 PF M 14 NA
7 7 7 16 1977 2 PE F NA NA
8 8 7 16 1977 1 DM M 37 NA
9 9 7 16 1977 1 DM F 34 NA
10 10 7 16 1977 6 PF F 20 NA
# ℹ 16,868 more rows
# ℹ 5 more variables: genus <chr>, species <chr>, taxa <chr>, plot_type <chr>,
# date <chr>Since our new column gets moved all the way to the end, it doesn’t
end up printing out. We can use the relocate() function to
put it after our year column:
R
surveys %>%
mutate(date = paste(year, month, day, sep = "-")) %>%
relocate(date, .after = year)
OUTPUT
# A tibble: 16,878 × 14
record_id month day year date plot_id species_id sex hindfoot_length
<dbl> <dbl> <dbl> <dbl> <chr> <dbl> <chr> <chr> <dbl>
1 1 7 16 1977 1977-7-… 2 NL M 32
2 2 7 16 1977 1977-7-… 3 NL M 33
3 3 7 16 1977 1977-7-… 2 DM F 37
4 4 7 16 1977 1977-7-… 7 DM M 36
5 5 7 16 1977 1977-7-… 3 DM M 35
6 6 7 16 1977 1977-7-… 1 PF M 14
7 7 7 16 1977 1977-7-… 2 PE F NA
8 8 7 16 1977 1977-7-… 1 DM M 37
9 9 7 16 1977 1977-7-… 1 DM F 34
10 10 7 16 1977 1977-7-… 6 PF F 20
# ℹ 16,868 more rows
# ℹ 5 more variables: weight <dbl>, genus <chr>, species <chr>, taxa <chr>,
# plot_type <chr>Now we can see that we have a character column that contains our date
string. However, it’s not truly a date column. Dates are a type of
numeric variable with a defined, ordered scale. To turn this column into
a proper date, we will use a function from the tidyverse’s
lubridate package, which has lots of useful functions for
working with dates. The function ymd() will parse a date
string that has the order year-month-day. Let’s load the package and use
ymd().
R
library(lubridate)
surveys %>%
mutate(date = paste(year, month, day, sep = "-"),
date = ymd(date)) %>%
relocate(date, .after = year)
OUTPUT
# A tibble: 16,878 × 14
record_id month day year date plot_id species_id sex
<dbl> <dbl> <dbl> <dbl> <date> <dbl> <chr> <chr>
1 1 7 16 1977 1977-07-16 2 NL M
2 2 7 16 1977 1977-07-16 3 NL M
3 3 7 16 1977 1977-07-16 2 DM F
4 4 7 16 1977 1977-07-16 7 DM M
5 5 7 16 1977 1977-07-16 3 DM M
6 6 7 16 1977 1977-07-16 1 PF M
7 7 7 16 1977 1977-07-16 2 PE F
8 8 7 16 1977 1977-07-16 1 DM M
9 9 7 16 1977 1977-07-16 1 DM F
10 10 7 16 1977 1977-07-16 6 PF F
# ℹ 16,868 more rows
# ℹ 6 more variables: hindfoot_length <dbl>, weight <dbl>, genus <chr>,
# species <chr>, taxa <chr>, plot_type <chr>R
surveys %>%
mutate(date = paste(year, month, day, sep = "-"),
date = as.Date(date)) %>%
relocate(date, .after = year)
OUTPUT
# A tibble: 16,878 × 14
record_id month day year date plot_id species_id sex
<dbl> <dbl> <dbl> <dbl> <date> <dbl> <chr> <chr>
1 1 7 16 1977 1977-07-16 2 NL M
2 2 7 16 1977 1977-07-16 3 NL M
3 3 7 16 1977 1977-07-16 2 DM F
4 4 7 16 1977 1977-07-16 7 DM M
5 5 7 16 1977 1977-07-16 3 DM M
6 6 7 16 1977 1977-07-16 1 PF M
7 7 7 16 1977 1977-07-16 2 PE F
8 8 7 16 1977 1977-07-16 1 DM M
9 9 7 16 1977 1977-07-16 1 DM F
10 10 7 16 1977 1977-07-16 6 PF F
# ℹ 16,868 more rows
# ℹ 6 more variables: hindfoot_length <dbl>, weight <dbl>, genus <chr>,
# species <chr>, taxa <chr>, plot_type <chr>Now we can see that our date column has the type
date as well. In this example, we created our column with
two separate lines in mutate(), but we can combine them
into one:
R
# using nested functions
surveys %>%
mutate(date = ymd(paste(year, month, day, sep = "-"))) %>%
relocate(date, .after = year)
OUTPUT
# A tibble: 16,878 × 14
record_id month day year date plot_id species_id sex
<dbl> <dbl> <dbl> <dbl> <date> <dbl> <chr> <chr>
1 1 7 16 1977 1977-07-16 2 NL M
2 2 7 16 1977 1977-07-16 3 NL M
3 3 7 16 1977 1977-07-16 2 DM F
4 4 7 16 1977 1977-07-16 7 DM M
5 5 7 16 1977 1977-07-16 3 DM M
6 6 7 16 1977 1977-07-16 1 PF M
7 7 7 16 1977 1977-07-16 2 PE F
8 8 7 16 1977 1977-07-16 1 DM M
9 9 7 16 1977 1977-07-16 1 DM F
10 10 7 16 1977 1977-07-16 6 PF F
# ℹ 16,868 more rows
# ℹ 6 more variables: hindfoot_length <dbl>, weight <dbl>, genus <chr>,
# species <chr>, taxa <chr>, plot_type <chr>R
# using a pipe *inside* mutate()
surveys %>%
mutate(date = paste(year, month, day,
sep = "-") %>% ymd()) %>%
relocate(date, .after = year)
OUTPUT
# A tibble: 16,878 × 14
record_id month day year date plot_id species_id sex
<dbl> <dbl> <dbl> <dbl> <date> <dbl> <chr> <chr>
1 1 7 16 1977 1977-07-16 2 NL M
2 2 7 16 1977 1977-07-16 3 NL M
3 3 7 16 1977 1977-07-16 2 DM F
4 4 7 16 1977 1977-07-16 7 DM M
5 5 7 16 1977 1977-07-16 3 DM M
6 6 7 16 1977 1977-07-16 1 PF M
7 7 7 16 1977 1977-07-16 2 PE F
8 8 7 16 1977 1977-07-16 1 DM M
9 9 7 16 1977 1977-07-16 1 DM F
10 10 7 16 1977 1977-07-16 6 PF F
# ℹ 16,868 more rows
# ℹ 6 more variables: hindfoot_length <dbl>, weight <dbl>, genus <chr>,
# species <chr>, taxa <chr>, plot_type <chr>R
surveys %>%
mutate(date = ymd(paste(year, month, day, sep = "-"))) %>%
ggplot(aes(x = date, y = weight)) +
geom_jitter(alpha = 0.1)
WARNING
Warning: Removed 1692 rows containing missing values or values outside the scale range
(`geom_point()`).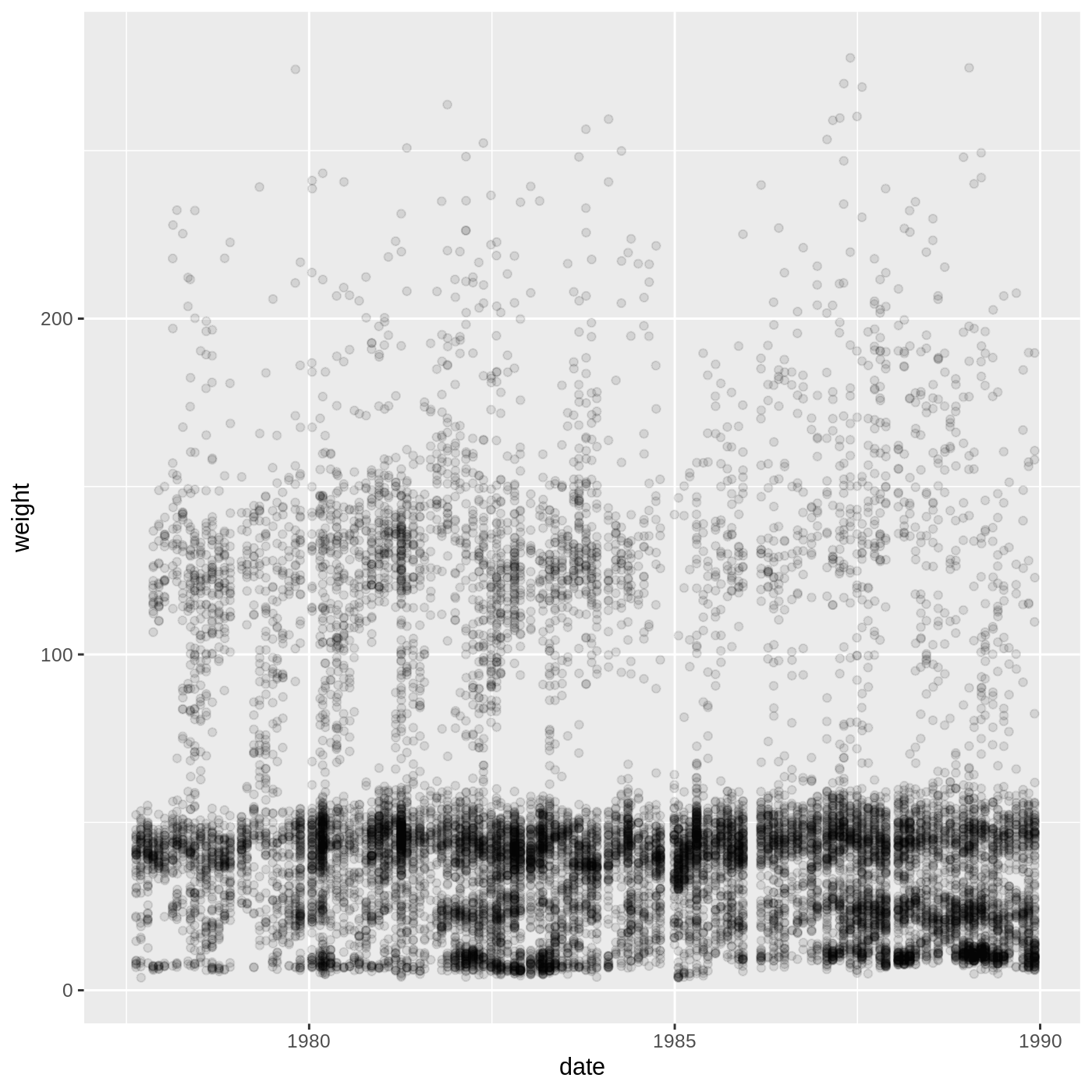
This isn’t necessarily the most useful plot, but we will learn some techniques that will help produce nice time series plots
The split-apply-combine approach
Many data analysis tasks can be achieved using the
split-apply-combine approach: you split the data into groups, apply some
analysis to each group, and combine the results in some way.
dplyr has a few convenient functions to enable this
approach, the main two being group_by() and
summarize().
group_by() takes a data.frame and the name of one or
more columns with categorical values that define the groups.
summarize() then collapses each group into a one-row
summary of the group, giving you back a data.frame with one row per
group. The syntax for summarize() is similar to
mutate(), where you define new columns based on values of
other columns. Let’s try calculating the mean weight of all our animals
by sex.
R
surveys %>%
group_by(sex) %>%
summarize(mean_weight = mean(weight, na.rm = T))
OUTPUT
# A tibble: 3 × 2
sex mean_weight
<chr> <dbl>
1 F 53.1
2 M 53.2
3 <NA> 74.0You can see that the mean weight for males is slightly higher than
for females, but that animals whose sex is unknown have much higher
weights. This is probably due to small sample size, but we should check
to be sure. Like mutate(), we can define multiple columns
in one summarize() call. The function n() will
count the number of rows in each group.
R
surveys %>%
group_by(sex) %>%
summarize(mean_weight = mean(weight, na.rm = T),
n = n())
OUTPUT
# A tibble: 3 × 3
sex mean_weight n
<chr> <dbl> <int>
1 F 53.1 7318
2 M 53.2 8260
3 <NA> 74.0 1300You will often want to create groups based on multiple columns. For
example, we might be interested in the mean weight of every species +
sex combination. All we have to do is add another column to our
group_by() call.
R
surveys %>%
group_by(species_id, sex) %>%
summarize(mean_weight = mean(weight, na.rm = T),
n = n())
OUTPUT
`summarise()` has grouped output by 'species_id'. You can override using the
`.groups` argument.OUTPUT
# A tibble: 67 × 4
# Groups: species_id [36]
species_id sex mean_weight n
<chr> <chr> <dbl> <int>
1 AB <NA> NaN 223
2 AH <NA> NaN 136
3 BA M 7 3
4 CB <NA> NaN 23
5 CM <NA> NaN 13
6 CQ <NA> NaN 16
7 CS <NA> NaN 1
8 CV <NA> NaN 1
9 DM F 40.7 2522
10 DM M 44.0 3108
# ℹ 57 more rowsOur resulting data.frame is much larger, since we have a greater
number of groups. We also see a strange value showing up in our
mean_weight column: NaN. This stands for “Not
a Number”, and it often results from trying to do an operation a vector
with zero entries. How can a vector have zero entries? Well, if a
particular group (like the AB species ID + NA sex group)
has only NA values for weight, then the
na.rm = T argument in mean() will remove
all the values prior to calculating the mean. The
result will be a value of NaN. Since we are not
particularly interested in these values, let’s add a step to our
pipeline to remove rows where weight is NA
before doing any other steps. This means that any
groups with only NA values will disappear from our
data.frame before we formally create the groups with
group_by().
R
surveys %>%
filter(!is.na(weight)) %>%
group_by(species_id, sex) %>%
summarize(mean_weight = mean(weight),
n = n())
OUTPUT
`summarise()` has grouped output by 'species_id'. You can override using the
`.groups` argument.OUTPUT
# A tibble: 46 × 4
# Groups: species_id [18]
species_id sex mean_weight n
<chr> <chr> <dbl> <int>
1 BA M 7 3
2 DM F 40.7 2460
3 DM M 44.0 3013
4 DM <NA> 37 8
5 DO F 48.4 679
6 DO M 49.3 748
7 DO <NA> 44 1
8 DS F 118. 1055
9 DS M 123. 1184
10 DS <NA> 121. 16
# ℹ 36 more rowsThat looks better! It’s often useful to take a look at the results in
some order, like the lowest mean weight to highest. We can use the
arrange() function for that:
R
surveys %>%
filter(!is.na(weight)) %>%
group_by(species_id, sex) %>%
summarize(mean_weight = mean(weight),
n = n()) %>%
arrange(mean_weight)
OUTPUT
`summarise()` has grouped output by 'species_id'. You can override using the
`.groups` argument.OUTPUT
# A tibble: 46 × 4
# Groups: species_id [18]
species_id sex mean_weight n
<chr> <chr> <dbl> <int>
1 PF <NA> 6 2
2 BA M 7 3
3 PF F 7.09 215
4 PF M 7.10 296
5 RM M 9.92 678
6 RM <NA> 10.4 7
7 RM F 10.7 629
8 RF M 12.4 16
9 RF F 13.7 46
10 PP <NA> 15 2
# ℹ 36 more rowsIf we want to reverse the order, we can wrap the column name in
desc():
R
surveys %>%
filter(!is.na(weight)) %>%
group_by(species_id, sex) %>%
summarize(mean_weight = mean(weight),
n = n()) %>%
arrange(desc(mean_weight))
OUTPUT
`summarise()` has grouped output by 'species_id'. You can override using the
`.groups` argument.OUTPUT
# A tibble: 46 × 4
# Groups: species_id [18]
species_id sex mean_weight n
<chr> <chr> <dbl> <int>
1 NL M 168. 355
2 NL <NA> 164. 9
3 NL F 151. 460
4 SS M 130 1
5 DS M 123. 1184
6 DS <NA> 121. 16
7 DS F 118. 1055
8 SH F 79.2 61
9 SH M 67.6 34
10 SF F 58.3 3
# ℹ 36 more rowsYou may have seen several messages saying
summarise() has grouped output by 'species_id'. You can override using the .groups argument.
These are warning you that your resulting data.frame has retained some
group structure, which means any subsequent operations on that
data.frame will happen at the group level. If you look at the resulting
data.frame printed out in your console, you will see these lines:
# A tibble: 46 × 4
# Groups: species_id [18]They tell us we have a data.frame with 46 rows, 4 columns, and a
group variable species_id, for which there are 18 groups.
We will see something similar if we use group_by()
alone:
R
surveys %>%
group_by(species_id, sex)
OUTPUT
# A tibble: 16,878 × 13
# Groups: species_id, sex [67]
record_id month day year plot_id species_id sex hindfoot_length weight
<dbl> <dbl> <dbl> <dbl> <dbl> <chr> <chr> <dbl> <dbl>
1 1 7 16 1977 2 NL M 32 NA
2 2 7 16 1977 3 NL M 33 NA
3 3 7 16 1977 2 DM F 37 NA
4 4 7 16 1977 7 DM M 36 NA
5 5 7 16 1977 3 DM M 35 NA
6 6 7 16 1977 1 PF M 14 NA
7 7 7 16 1977 2 PE F NA NA
8 8 7 16 1977 1 DM M 37 NA
9 9 7 16 1977 1 DM F 34 NA
10 10 7 16 1977 6 PF F 20 NA
# ℹ 16,868 more rows
# ℹ 4 more variables: genus <chr>, species <chr>, taxa <chr>, plot_type <chr>What we get back is the entire surveys data.frame, but
with the grouping variables added: 67 groups of species_id
+ sex combinations. Groups are often maintained throughout
a pipeline, and if you assign the resulting data.frame to a new object,
it will also have those groups. This can lead to confusing results if
you forget about the grouping and want to carry out operations on the
whole data.frame, not by group. Therefore, it is a good habit to remove
the groups at the end of a pipeline containing
group_by():
R
surveys %>%
filter(!is.na(weight)) %>%
group_by(species_id, sex) %>%
summarize(mean_weight = mean(weight),
n = n()) %>%
arrange(desc(mean_weight)) %>%
ungroup()
OUTPUT
`summarise()` has grouped output by 'species_id'. You can override using the
`.groups` argument.OUTPUT
# A tibble: 46 × 4
species_id sex mean_weight n
<chr> <chr> <dbl> <int>
1 NL M 168. 355
2 NL <NA> 164. 9
3 NL F 151. 460
4 SS M 130 1
5 DS M 123. 1184
6 DS <NA> 121. 16
7 DS F 118. 1055
8 SH F 79.2 61
9 SH M 67.6 34
10 SF F 58.3 3
# ℹ 36 more rowsNow our data.frame just says # A tibble: 46 × 4 at the
top, with no groups.
While it is common that you will want to get the one-row-per-group
summary that summarise() provides, there are times where
you want to calculate a per-group value but keep all the rows in your
data.frame. For example, we might want to know the mean weight for each
species ID + sex combination, and then we might want to know how far
from that mean value each observation in the group is. For this, we can
use group_by() and mutate() together:
R
surveys %>%
filter(!is.na(weight)) %>%
group_by(species_id, sex) %>%
mutate(mean_weight = mean(weight),
weight_diff = weight - mean_weight)
OUTPUT
# A tibble: 15,186 × 15
# Groups: species_id, sex [46]
record_id month day year plot_id species_id sex hindfoot_length weight
<dbl> <dbl> <dbl> <dbl> <dbl> <chr> <chr> <dbl> <dbl>
1 63 8 19 1977 3 DM M 35 40
2 64 8 19 1977 7 DM M 37 48
3 65 8 19 1977 4 DM F 34 29
4 66 8 19 1977 4 DM F 35 46
5 67 8 19 1977 7 DM M 35 36
6 68 8 19 1977 8 DO F 32 52
7 69 8 19 1977 2 PF M 15 8
8 70 8 19 1977 3 OX F 21 22
9 71 8 19 1977 7 DM F 36 35
10 74 8 19 1977 8 PF M 12 7
# ℹ 15,176 more rows
# ℹ 6 more variables: genus <chr>, species <chr>, taxa <chr>, plot_type <chr>,
# mean_weight <dbl>, weight_diff <dbl>Since we get all our columns back, the new columns are at the very
end and don’t print out in the console. Let’s use select()
to just look at the columns of interest. Inside select() we
can use the contains() function to get any column
containing the word “weight” in the name:
R
surveys %>%
filter(!is.na(weight)) %>%
group_by(species_id, sex) %>%
mutate(mean_weight = mean(weight),
weight_diff = weight - mean_weight) %>%
select(species_id, sex, contains("weight"))
OUTPUT
# A tibble: 15,186 × 5
# Groups: species_id, sex [46]
species_id sex weight mean_weight weight_diff
<chr> <chr> <dbl> <dbl> <dbl>
1 DM M 40 44.0 -4.00
2 DM M 48 44.0 4.00
3 DM F 29 40.7 -11.7
4 DM F 46 40.7 5.28
5 DM M 36 44.0 -8.00
6 DO F 52 48.4 3.63
7 PF M 8 7.10 0.902
8 OX F 22 21 1
9 DM F 35 40.7 -5.72
10 PF M 7 7.10 -0.0980
# ℹ 15,176 more rowsWhat happens with the group_by() + mutate()
combination is similar to using summarize(): for each
group, the mean weight is calculated. However, instead of reporting only
one row per group, the mean weight for each group is added to each row
in that group. For each row in a group (like DM species ID + M sex), you
will see the same value in mean_weight.
R
surveys_daily_counts <- surveys %>%
mutate(date = ymd(paste(year, month, day, sep = "-"))) %>%
group_by(date, sex) %>%
summarize(n = n())
OUTPUT
`summarise()` has grouped output by 'date'. You can override using the
`.groups` argument.R
# shorter approach using count()
surveys_daily_counts <- surveys %>%
mutate(date = ymd(paste(year, month, day, sep = "-"))) %>%
count(date, sex)
Challenge 4: Making a time series (continued)
- Now use the data.frame you just made to plot the daily number of
animals of each sex caught over time. It’s up to you what
geomto use, but alineplot might be a good choice. You should also think about how to differentiate which data corresponds to which sex.
R
surveys_daily_counts %>%
ggplot(aes(x = date, y = n, color = sex)) +
geom_line()
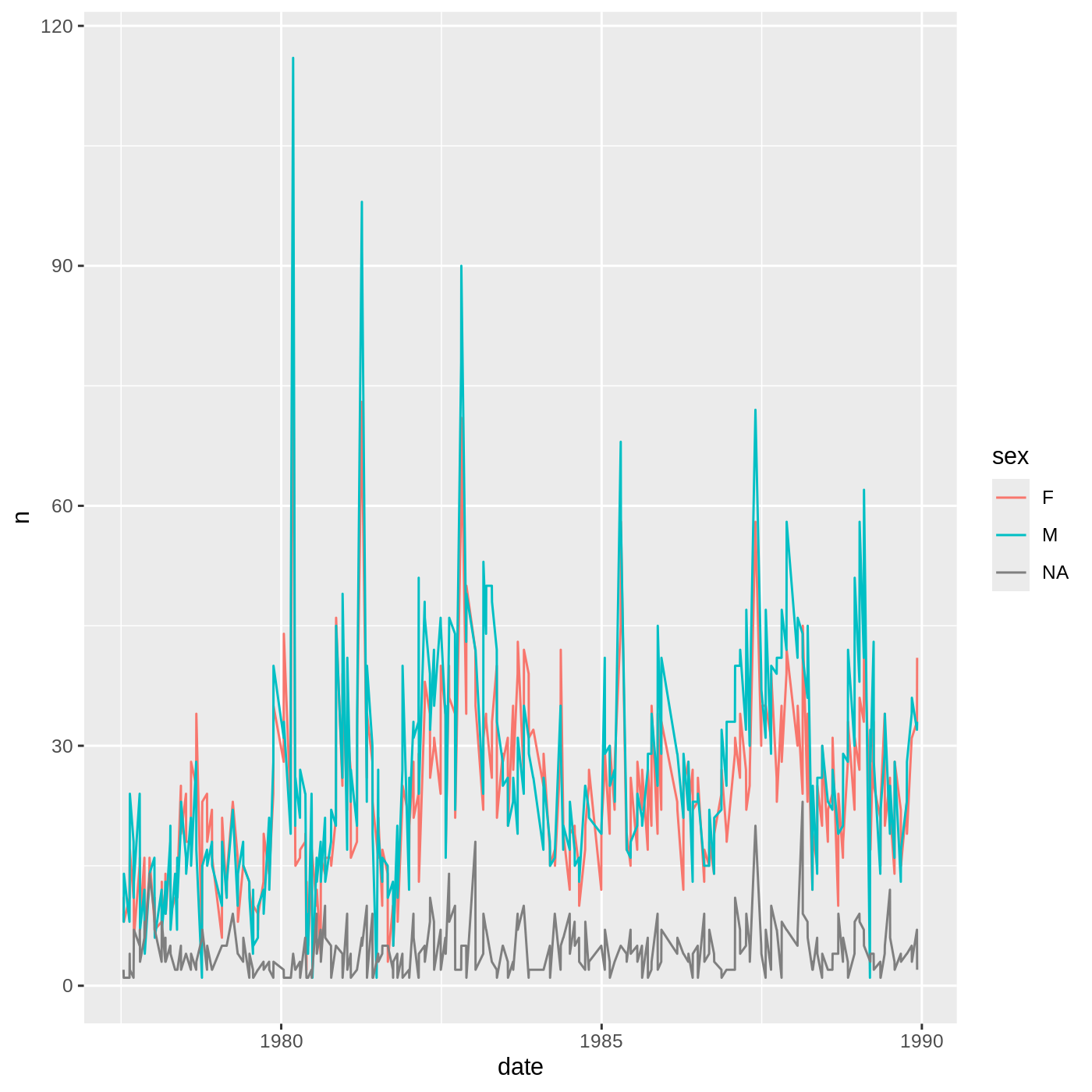
Reshaping data with tidyr
Let’s say we are interested in comparing the mean weights of each
species across our different plots. We can begin this process using the
group_by() + summarize() approach:
R
sp_by_plot <- surveys %>%
filter(!is.na(weight)) %>%
group_by(species_id, plot_id) %>%
summarise(mean_weight = mean(weight)) %>%
arrange(species_id, plot_id)
OUTPUT
`summarise()` has grouped output by 'species_id'. You can override using the
`.groups` argument.R
sp_by_plot
OUTPUT
# A tibble: 300 × 3
# Groups: species_id [18]
species_id plot_id mean_weight
<chr> <dbl> <dbl>
1 BA 3 8
2 BA 21 6.5
3 DM 1 42.7
4 DM 2 42.6
5 DM 3 41.2
6 DM 4 41.9
7 DM 5 42.6
8 DM 6 42.1
9 DM 7 43.2
10 DM 8 43.4
# ℹ 290 more rowsThat looks great, but it is a bit difficult to compare values across
plots. It would be nice if we could reshape this data.frame to make
those comparisons easier. Well, the tidyr package from the
tidyverse has a pair of functions that allow you to reshape
data by pivoting it: pivot_wider() and
pivot_longer(). pivot_wider() will make the
data wider, which means increasing the number of columns and reducing
the number of rows. pivot_longer() will do the opposite,
reducing the number of columns and increasing the number of rows.
In this case, it might be nice to create a data.frame where each
species has its own row, and each plot has its own column containing the
mean weight for a given species. We will use pivot_wider()
to reshape our data in this way. It takes 3 arguments:
- the name of the data.frame
-
names_from: which column should be used to generate the names of the new columns? -
values_from: which column should be used to fill in the values of the new columns?
Any columns not used for names_from or
values_from will not be pivoted.
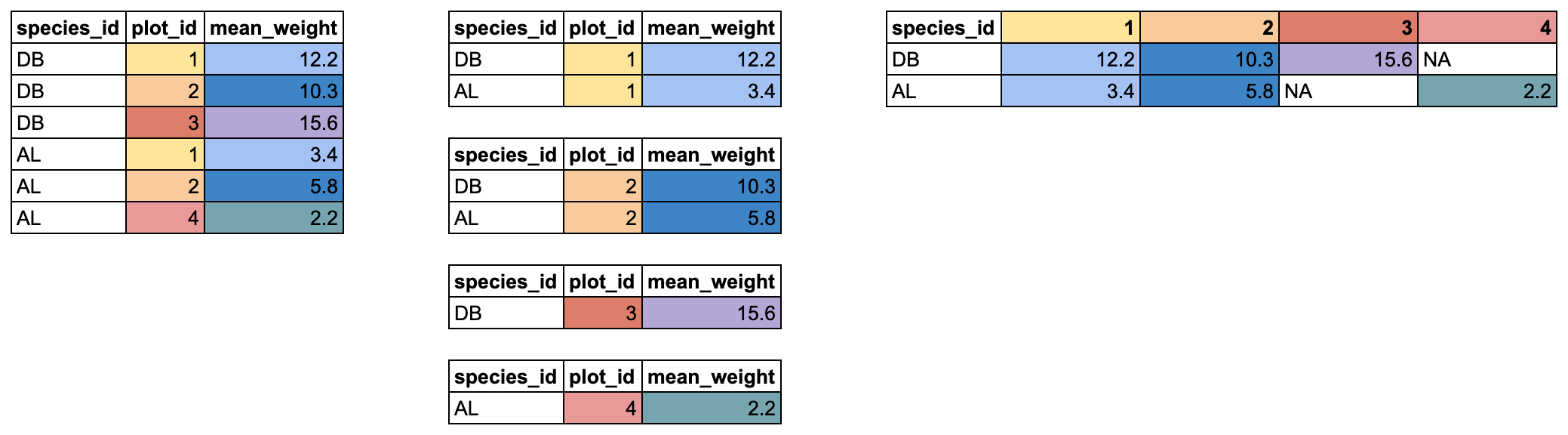
In our case, we want the new columns to be named from our
plot_id column, with the values coming from the
mean_weight column. We can pipe our data.frame right into
pivot_wider() and add those two arguments:
R
sp_by_plot_wide <- sp_by_plot %>%
pivot_wider(names_from = plot_id,
values_from = mean_weight)
sp_by_plot_wide
OUTPUT
# A tibble: 18 × 25
# Groups: species_id [18]
species_id `3` `21` `1` `2` `4` `5` `6` `7` `8`
<chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
1 BA 8 6.5 NA NA NA NA NA NA NA
2 DM 41.2 41.5 42.7 42.6 41.9 42.6 42.1 43.2 43.4
3 DO 42.7 NA 50.1 50.3 46.8 50.4 49.0 52 49.2
4 DS 128. NA 129. 125. 118. 111. 114. 126. 128.
5 NL 171. 136. 154. 171. 164. 192. 176. 170. 134.
6 OL 32.1 28.6 35.5 34 33.0 32.6 31.8 NA 30.3
7 OT 24.1 24.1 23.7 24.9 26.5 23.6 23.5 22 24.1
8 OX 22 NA NA 22 NA 20 NA NA NA
9 PE 22.7 19.6 21.6 22.0 NA 21 21.6 22.8 19.4
10 PF 7.12 7.23 6.57 6.89 6.75 7.5 7.54 7 6.78
11 PH 28 31 NA NA NA 29 NA NA NA
12 PM 20.1 23.6 23.7 23.9 NA 23.7 22.3 23.4 23
13 PP 17.1 13.6 14.3 16.4 14.8 19.8 16.8 NA 13.9
14 RF 14.8 17 NA 16 NA 14 12.1 13 NA
15 RM 10.3 9.89 10.9 10.6 10.4 10.8 10.6 10.7 9
16 SF NA 49 NA NA NA NA NA NA NA
17 SH 76.0 79.9 NA 88 NA 82.7 NA NA NA
18 SS NA NA NA NA NA NA NA NA NA
# ℹ 15 more variables: `9` <dbl>, `10` <dbl>, `11` <dbl>, `12` <dbl>,
# `13` <dbl>, `14` <dbl>, `15` <dbl>, `16` <dbl>, `17` <dbl>, `18` <dbl>,
# `19` <dbl>, `20` <dbl>, `22` <dbl>, `23` <dbl>, `24` <dbl>Now we’ve got our reshaped data.frame. There are a few things to
notice. First, we have a new column for each plot_id value.
There is one old column left in the data.frame: species_id.
It wasn’t used in pivot_wider(), so it stays, and now
contains a single entry for each unique species_id
value.
Finally, a lot of NAs have appeared. Some species aren’t
found in every plot, but because a data.frame has to have a value in
every row and every column, an NA is inserted. We can
double-check this to verify what is going on.
Looking in our new pivoted data.frame, we can see that there is an
NA value for the species BA in plot
1. Let’s take our sp_by_plot data.frame and
look for the mean_weight of that species + plot
combination.
R
sp_by_plot %>%
filter(species_id == "BA" & plot_id == 1)
OUTPUT
# A tibble: 0 × 3
# Groups: species_id [0]
# ℹ 3 variables: species_id <chr>, plot_id <dbl>, mean_weight <dbl>We get back 0 rows. There is no mean_weight for the
species BA in plot 1. This either happened
because no BA were ever caught in plot 1, or
because every BA caught in plot 1 had an
NA weight value and all the rows got removed when we used
filter(!is.na(weight)) in the process of making
sp_by_plot. Because there are no rows with that species +
plot combination, in our pivoted data.frame, the value gets filled with
NA.
There is another pivot_ function that does the opposite,
moving data from a wide to long format, called
pivot_longer(). It takes 3 arguments: cols for
the columns you want to pivot, names_to for the name of the
new column which will contain the old column names, and
values_to for the name of the new column which will contain
the old values.
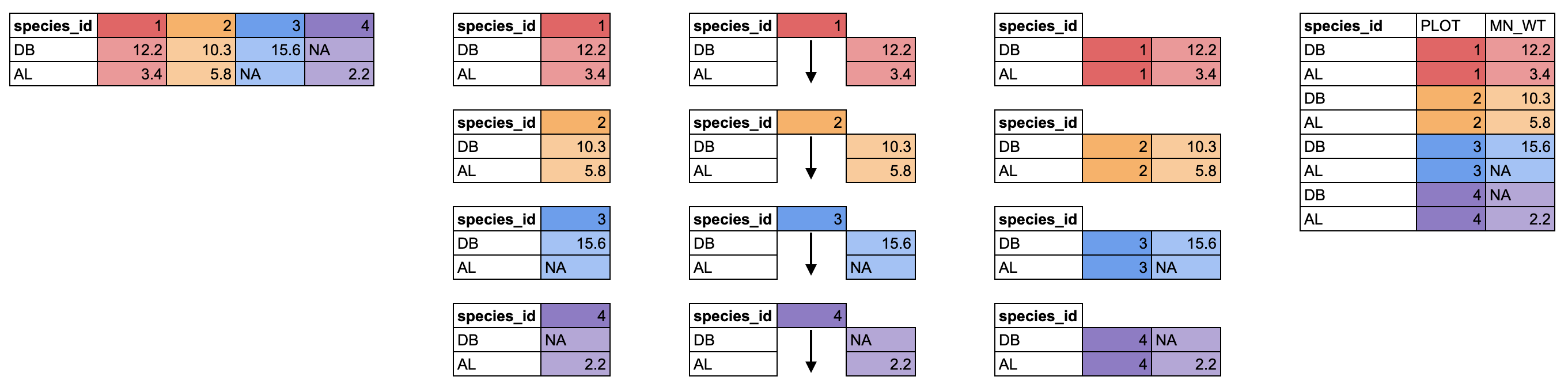
We can pivot our new wide data.frame to a long format using
pivot_longer(). We want to pivot all the columns except
species_id, and we will use PLOT for the new
column of plot IDs, and MEAN_WT for the new column of mean
weight values.
R
sp_by_plot_wide %>%
pivot_longer(cols = -species_id, names_to = "PLOT", values_to = "MEAN_WT")
OUTPUT
# A tibble: 432 × 3
# Groups: species_id [18]
species_id PLOT MEAN_WT
<chr> <chr> <dbl>
1 BA 3 8
2 BA 21 6.5
3 BA 1 NA
4 BA 2 NA
5 BA 4 NA
6 BA 5 NA
7 BA 6 NA
8 BA 7 NA
9 BA 8 NA
10 BA 9 NA
# ℹ 422 more rowsOne thing you will notice is that all those NA values
that got generated when we pivoted wider. However, we can filter those
out, which gets us back to the same data as sp_by_plot,
before we pivoted it wider.
R
sp_by_plot_wide %>%
pivot_longer(cols = -species_id, names_to = "PLOT", values_to = "MEAN_WT") %>%
filter(!is.na(MEAN_WT))
OUTPUT
# A tibble: 300 × 3
# Groups: species_id [18]
species_id PLOT MEAN_WT
<chr> <chr> <dbl>
1 BA 3 8
2 BA 21 6.5
3 DM 3 41.2
4 DM 21 41.5
5 DM 1 42.7
6 DM 2 42.6
7 DM 4 41.9
8 DM 5 42.6
9 DM 6 42.1
10 DM 7 43.2
# ℹ 290 more rowsData are often recorded in spreadsheets in a wider format, but lots
of tidyverse tools, especially ggplot2, like
data in a longer format, so pivot_longer() is often very
useful.
Exporting data
Let’s say we want to send the wide version of our
sb_by_plot data.frame to a colleague who doesn’t use R. In
this case, we might want to save it as a CSV file.
First, we might want to modify the names of the columns, since right
now they are bare numbers, which aren’t very informative. Luckily,
pivot_wider() has an argument names_prefix
which will allow us to add “plot_” to the start of each column.
R
sp_by_plot %>%
pivot_wider(names_from = plot_id, values_from = mean_weight,
names_prefix = "plot_")
OUTPUT
# A tibble: 18 × 25
# Groups: species_id [18]
species_id plot_3 plot_21 plot_1 plot_2 plot_4 plot_5 plot_6 plot_7 plot_8
<chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
1 BA 8 6.5 NA NA NA NA NA NA NA
2 DM 41.2 41.5 42.7 42.6 41.9 42.6 42.1 43.2 43.4
3 DO 42.7 NA 50.1 50.3 46.8 50.4 49.0 52 49.2
4 DS 128. NA 129. 125. 118. 111. 114. 126. 128.
5 NL 171. 136. 154. 171. 164. 192. 176. 170. 134.
6 OL 32.1 28.6 35.5 34 33.0 32.6 31.8 NA 30.3
7 OT 24.1 24.1 23.7 24.9 26.5 23.6 23.5 22 24.1
8 OX 22 NA NA 22 NA 20 NA NA NA
9 PE 22.7 19.6 21.6 22.0 NA 21 21.6 22.8 19.4
10 PF 7.12 7.23 6.57 6.89 6.75 7.5 7.54 7 6.78
11 PH 28 31 NA NA NA 29 NA NA NA
12 PM 20.1 23.6 23.7 23.9 NA 23.7 22.3 23.4 23
13 PP 17.1 13.6 14.3 16.4 14.8 19.8 16.8 NA 13.9
14 RF 14.8 17 NA 16 NA 14 12.1 13 NA
15 RM 10.3 9.89 10.9 10.6 10.4 10.8 10.6 10.7 9
16 SF NA 49 NA NA NA NA NA NA NA
17 SH 76.0 79.9 NA 88 NA 82.7 NA NA NA
18 SS NA NA NA NA NA NA NA NA NA
# ℹ 15 more variables: plot_9 <dbl>, plot_10 <dbl>, plot_11 <dbl>,
# plot_12 <dbl>, plot_13 <dbl>, plot_14 <dbl>, plot_15 <dbl>, plot_16 <dbl>,
# plot_17 <dbl>, plot_18 <dbl>, plot_19 <dbl>, plot_20 <dbl>, plot_22 <dbl>,
# plot_23 <dbl>, plot_24 <dbl>That looks better! Let’s save this data.frame as a new object.
R
surveys_sp <- sp_by_plot %>%
pivot_wider(names_from = plot_id, values_from = mean_weight,
names_prefix = "plot_")
surveys_sp
OUTPUT
# A tibble: 18 × 25
# Groups: species_id [18]
species_id plot_3 plot_21 plot_1 plot_2 plot_4 plot_5 plot_6 plot_7 plot_8
<chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
1 BA 8 6.5 NA NA NA NA NA NA NA
2 DM 41.2 41.5 42.7 42.6 41.9 42.6 42.1 43.2 43.4
3 DO 42.7 NA 50.1 50.3 46.8 50.4 49.0 52 49.2
4 DS 128. NA 129. 125. 118. 111. 114. 126. 128.
5 NL 171. 136. 154. 171. 164. 192. 176. 170. 134.
6 OL 32.1 28.6 35.5 34 33.0 32.6 31.8 NA 30.3
7 OT 24.1 24.1 23.7 24.9 26.5 23.6 23.5 22 24.1
8 OX 22 NA NA 22 NA 20 NA NA NA
9 PE 22.7 19.6 21.6 22.0 NA 21 21.6 22.8 19.4
10 PF 7.12 7.23 6.57 6.89 6.75 7.5 7.54 7 6.78
11 PH 28 31 NA NA NA 29 NA NA NA
12 PM 20.1 23.6 23.7 23.9 NA 23.7 22.3 23.4 23
13 PP 17.1 13.6 14.3 16.4 14.8 19.8 16.8 NA 13.9
14 RF 14.8 17 NA 16 NA 14 12.1 13 NA
15 RM 10.3 9.89 10.9 10.6 10.4 10.8 10.6 10.7 9
16 SF NA 49 NA NA NA NA NA NA NA
17 SH 76.0 79.9 NA 88 NA 82.7 NA NA NA
18 SS NA NA NA NA NA NA NA NA NA
# ℹ 15 more variables: plot_9 <dbl>, plot_10 <dbl>, plot_11 <dbl>,
# plot_12 <dbl>, plot_13 <dbl>, plot_14 <dbl>, plot_15 <dbl>, plot_16 <dbl>,
# plot_17 <dbl>, plot_18 <dbl>, plot_19 <dbl>, plot_20 <dbl>, plot_22 <dbl>,
# plot_23 <dbl>, plot_24 <dbl>Now we can save this data.frame to a CSV using the
write_csv() function from the readr package.
The first argument is the name of the data.frame, and the second is the
path to the new file we want to create, including the file extension
.csv.
R
write_csv(surveys_sp, "data/cleaned/surveys_meanweight_species_plot.csv")
If we go look into our data/cleaned_data folder, we will
see this new CSV file.
Key Points
- use
filter()to subset rows andselect()to subset columns - build up pipelines one step at a time before assigning the result
- it is often best to keep components of dates separate until needed,
then use
mutate()to make a date column -
group_by()can be used withsummarize()to collapse rows ormutate()to keep the same number of rows -
pivot_wider()andpivot_longer()are powerful for reshaping data, but you should plan out how to use them thoughtfully
